
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 6,368,838 – 6,368,931 |
| Length | 93 |
| Max. P | 0.530281 |

| Location | 6,368,838 – 6,368,931 |
|---|---|
| Length | 93 |
| Sequences | 13 |
| Columns | 108 |
| Reading direction | reverse |
| Mean pairwise identity | 67.23 |
| Shannon entropy | 0.68196 |
| G+C content | 0.42916 |
| Mean single sequence MFE | -21.80 |
| Consensus MFE | -9.92 |
| Energy contribution | -9.75 |
| Covariance contribution | -0.17 |
| Combinations/Pair | 1.50 |
| Mean z-score | -0.70 |
| Structure conservation index | 0.45 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.07 |
| SVM RNA-class probability | 0.530281 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 6368838 93 - 21146708 UCGCUCUUCUUCACCAGGAAGUUUAUGAAUUCGGGCGAGAAUCGAAAGCCCAUCUGGGUGAAGGCUGC-CAACAAGUUUGGUAAUUAGUGUAAA-------------- .((((..((((((((.(((.......(....)((((...........)))).))).)))))))).(((-(((.....))))))...))))....-------------- ( -26.60, z-score = -1.31, R) >droGri2.scaffold_15245 5192351 82 - 18325388 UCACUCUUUUUCACCAGAAAGUUGAUAAACUCGGGCGUAAAGCGAAAGCCCAUUUGAGUGAAAGCUGUUACAUAAGAAAAGC-------------------------- ........((((((((((.((((....)))).((((...........)))).)))).))))))(((.............)))-------------------------- ( -17.32, z-score = -0.98, R) >droVir3.scaffold_12875 16334828 86 - 20611582 UCACUCUUCUUUACCAGAAAGUUUAUGAACUCCGGCGUGAAGCGAAAGCCCAUUUGAGUAAAUGCUACACAAGAAAAAAGGUAAAU---------------------- .........((((((......((((((........))))))((....))...((((.(((.....))).))))......)))))).---------------------- ( -14.10, z-score = -0.26, R) >droMoj3.scaffold_6496 19444261 86 + 26866924 UCACUUUUCUUCACCAGGAAGUUUAUAAAUUCCGGCGUAAAGCGAAAGCCCAUUUGUGUGAAUGCUGUAAAAGAA----ACAUAUUACAU------------------ .....((((((.....((((.........))))(((((...((....)).(((....))).)))))....)))))----)..........------------------ ( -14.70, z-score = -0.57, R) >droWil1.scaffold_180569 261569 85 - 1405142 UCGCUCUUUUUCACCAGGAAGUUGAUAAACUCGGGCGUAAAACGGAAACCCAUUUGGGUAAAAGCUGCGAUAAAAAAUAGUUAGU----------------------- ((((.(((((...(((((.((((....)))).(((((.....))....))).)))))..)))))..))))...............----------------------- ( -17.50, z-score = -0.36, R) >droPer1.super_4 527891 89 - 7162766 UCGCUCUUCUUCACCAGAAAGUUGAUGAAUUCCGGCGAAAACCGAAAGCCCAUCUGAGUGAAGGCUGUGGGCAAGGAAAAGCCAUUAGU------------------- .(((...(((((((((((..((((........))))......(....)....)))).)))))))..)))(((........)))......------------------- ( -23.10, z-score = -0.44, R) >dp4.chr3 3542383 89 + 19779522 UCGCUCUUCUUCACCAGAAAGUUGAUGAAUUCCGGCGAAAACCGAAAGCCCAUCUGAGUGAAGGCUGUGGGCAAGGAAAAGCCAUUAGU------------------- .(((...(((((((((((..((((........))))......(....)....)))).)))))))..)))(((........)))......------------------- ( -23.10, z-score = -0.44, R) >droAna3.scaffold_13266 17841418 97 - 19884421 UCGCUCUUCUUCACCAAGAAGCCAAUAAACUCUGGAGUGAAGCGGAAACCCAUUUGGGUGAAGGCUAUGGGGAAAUUAUAUAUUUUGCAUUCGUUAA----------- (((((((((((....))))))(((........))))))))((((((.((((....))))..........(.(((((.....))))).).))))))..----------- ( -21.50, z-score = 0.08, R) >droEre2.scaffold_4845 3205501 89 + 22589142 UCGCUCUUCUUCACCAGAAAGUUGAUGAACUCGGGCGAGAAUCGAAAGCCCAUCUGGGUGAAGGCUGCAAAACAAGUGUGAUAAUUAUU------------------- ((((....((((((((((.((((....)))).((((...........)))).))).)))))))(((........)))))))........------------------- ( -25.00, z-score = -1.50, R) >droYak2.chr2R 18367827 102 + 21139217 UCGCUCUUCUUCACCAGGAAGUUAAUGAAGUCGGGCGAGAAUCGAAAGCCCAUCUGGGUGAAGGCUGC-AAGCAAGUGUGG-GAUUAUAGCGAAGGCCCUCACA---- ..((...((((((((.(((.......(....)((((...........)))).))).))))))))..))-......(((.((-(.((......))..))).))).---- ( -30.80, z-score = -0.21, R) >droSec1.super_1 3980428 93 - 14215200 UCGCUCUUCUUCACCAGGAAGUUGAUGAAUUCGGGCGAGAAUCGAAAGCCCAUCUGGGUGAAGGCUGC-GAACAAAUUUGGUAAUUAGUGUAAA-------------- ((((...((((((((.(((....((.....))((((...........)))).))).))))))))..))-)).......................-------------- ( -27.30, z-score = -1.50, R) >droSim1.chr2R 4997377 93 - 19596830 UCGCUCUUCUUCACCAGGAAGUUGAUGAACUCGGGCGAGAAUCGAAAGCCCAUCUGGGUGAAGGCUGC-GAACAAGUUUGGUAAUUAGUGUAAA-------------- ((((...((((((((.(((((((....)))).((((...........)))).))).))))))))..))-)).......................-------------- ( -27.90, z-score = -1.32, R) >triCas2.ChLG1X 1416923 107 - 8109244 UCGCUUUUAGCAAUUAAAAAUUUUACGAAAUCGGGGGAAAAUUUAAAUCCCAUCUGUUGCAACGCUAA-AAAUAAAAUAAGAUCCAGCAAUAAUUCUAUUAACACCGA ..((.....))...................((((((((.........))))...((((((....((..-..........)).....))))))............)))) ( -14.50, z-score = -0.27, R) >consensus UCGCUCUUCUUCACCAGGAAGUUGAUGAACUCGGGCGAGAAUCGAAAGCCCAUCUGGGUGAAGGCUGC_AAACAAGAAAGGUAAUUAGU___________________ .......(((((((((((.((((....))))..(((...........)))..)))).)))))))............................................ ( -9.92 = -9.75 + -0.17)


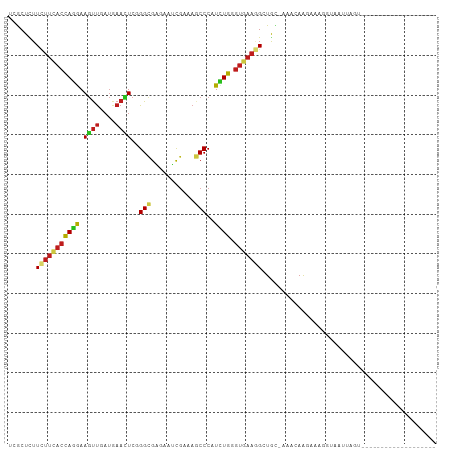
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:12:27 2011