
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 6,340,633 – 6,340,730 |
| Length | 97 |
| Max. P | 0.764260 |

| Location | 6,340,633 – 6,340,730 |
|---|---|
| Length | 97 |
| Sequences | 9 |
| Columns | 115 |
| Reading direction | forward |
| Mean pairwise identity | 56.96 |
| Shannon entropy | 0.86819 |
| G+C content | 0.45134 |
| Mean single sequence MFE | -25.70 |
| Consensus MFE | -7.43 |
| Energy contribution | -7.34 |
| Covariance contribution | -0.09 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.33 |
| Structure conservation index | 0.29 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.62 |
| SVM RNA-class probability | 0.764260 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 6340633 97 + 21146708 -----AAUUUGCCGCCUU-----UGACUUGACUGUUGUUACUUCAUUCGA--ACUGACAAUGGUGUUGCCCCAGGCAACGAAAUGCUCA-AUUAAAU-GCGAGAUUUAAGU---- -----.......(((.((-----(((.(((((((((((((.(((....))--).))))))))))((((((...))))))........))-)))))).-)))..........---- ( -26.00, z-score = -2.63, R) >droSim1.chr2R 4968983 102 + 19596830 -----AAUUUGCCGCCUUUGCUUUGACUUGACUGUUGUUACUUCAUUCGA--ACUGACAAUGGGGUUGCCACAGGCAACUGAAUGAUCA-AUUAAAU-GCGAGAUUUAAGU---- -----...........((((((((((.(((((((((((((.(((....))--).)))))))))(((((((...)))))))......)))-)))))).-)))))........---- ( -26.80, z-score = -2.10, R) >droSec1.super_1 3952337 102 + 14215200 -----AAUUUGCCGCCUUUGCUUUGACUUGACUGUUGUUACUUCAUUCGA--ACUGACAAUGGGGUUGCCACAGGCAACGAAAUGCUCA-AUUAAAU-GCGAGAUUUAAGU---- -----...........((((((((((.(((((((((((((.(((....))--).))))))))).((((((...)))))).......)))-)))))).-)))))........---- ( -25.60, z-score = -1.41, R) >droYak2.chr2L 18977896 96 + 22324452 -----AAUUUGCCGCCUU-----UGAGUUGACUGUUGUUCCUCCAUUCGC--ACUGACAAUGG-GUUGCCCCAGGCAACAAACUGCUCA-AUUAAAU-GCAAGAUUUAAGU---- -----..(((((.....(-----(((((.(.((((((((...........--...))))))))-((((((...))))))...).)))))-)......-)))))........---- ( -22.94, z-score = -0.76, R) >droEre2.scaffold_4929 18600742 96 - 26641161 -----AAUUUGCCGCCUU-----UGGCUUGGCUGUUGUUUCUCGAUUCGU--AGCUACAAUGG-GUUGCCACAGGCAACAAAAUGCUCA-AUUAAAU-GCGAGAUUUAAGU---- -----..((((..((((.-----((((..(.(((((((..((((...)).--))..)))))))-.).)))).))))..)))).(((...-.......-)))..........---- ( -23.50, z-score = -0.03, R) >droAna3.scaffold_13266 17810738 76 + 19884421 --------------------------------UGCCACACAGUGGCAGUG--ACUUCUCAUCGAGUAGCUCCGAGCCACAAUAUCUUCA-AUUAACUUGGGAGAUCUCAUU---- --------------------------------((((((...))))))(((--(..((((.((((((.((.....)).....((......-..))))))))))))..)))).---- ( -19.60, z-score = -1.58, R) >dp4.chr3 3515933 110 - 19779522 -----AAACCGCCGUGCUCUGCCCUGCCCGGCUGUUUGCCUGCCAUUCCGCUGCUCGUAGGAGGGUUGCCCCGGGCAACAGCAUACUCGUAUUAUAUUGAAUGACAGAUACCCCU -----((((.((((.((........)).)))).))))..(((.(((((.(((.(((....))).((((((...)))))))))(((....)))......))))).)))........ ( -31.60, z-score = 0.05, R) >droPer1.super_4 501357 107 + 7162766 -----AAACCGCCGUGCUCUGCCCUGCCCGGCUGUUUGCCUGCCAUUCCG---CUCGUAGGAGGGUUGCCCCGGGCAACAGCAUACUCGUAUUAUAUUGAAUGACAGAUCCCCCU -----((((.((((.((........)).)))).))))..(((.(((((..---...(((.(((.((((((...))))))......))).)))......))))).)))........ ( -32.00, z-score = -0.48, R) >droGri2.scaffold_15245 5159827 92 + 18325388 AGAAGUGUAUGUCAUCUUCGGUGUGCCUUUGAUAGCUUUUUUUUGUUUGU---UUUUUGUUUGUGUUGCCCCAGGCAACACAAU--UCA-AUUAAAU-G---------------- .((((.((((..(......)..))))))))..............(((((.---...(((.((((((((((...)))))))))).--.))-).)))))-.---------------- ( -23.30, z-score = -2.99, R) >consensus _____AAUUUGCCGCCUU_____UGACUUGACUGUUGUUCCUUCAUUCGA__ACUGACAAUGGGGUUGCCCCAGGCAACAAAAUGCUCA_AUUAAAU_GCGAGAUUUAAGU____ ................................................................((((((...)))))).................................... ( -7.43 = -7.34 + -0.09)
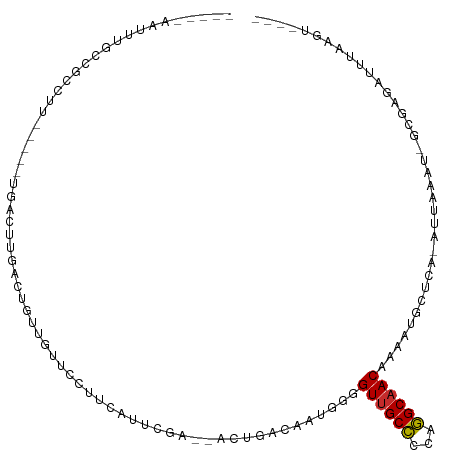


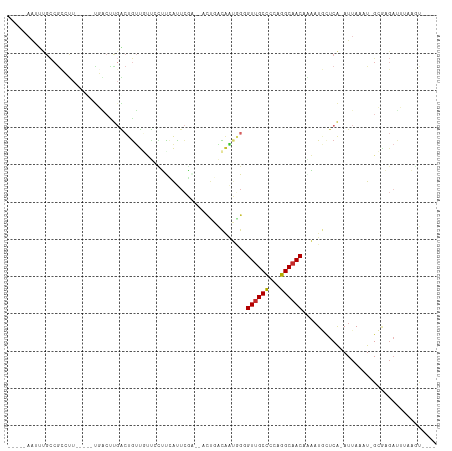
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:12:22 2011