
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 6,267,836 – 6,267,926 |
| Length | 90 |
| Max. P | 0.500000 |

| Location | 6,267,836 – 6,267,926 |
|---|---|
| Length | 90 |
| Sequences | 8 |
| Columns | 116 |
| Reading direction | forward |
| Mean pairwise identity | 63.37 |
| Shannon entropy | 0.63367 |
| G+C content | 0.44111 |
| Mean single sequence MFE | -21.07 |
| Consensus MFE | -7.10 |
| Energy contribution | -7.29 |
| Covariance contribution | 0.19 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.48 |
| Structure conservation index | 0.34 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.01 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
>dm3.chr2R 6267836 90 + 21146708 ------GGCAGCCGGAGGAAGCAGAGCAUUCCAAGCGCAUUUAAUGCGCAUAAUGUCAAAUGCAACAAAGCAACGGA----AACAACAAACCUCAGUACA---------------- ------........((((..(((..(((((....((((((...))))))..)))))....)))...........(..----..)......))))......---------------- ( -21.50, z-score = -1.58, R) >droSec1.super_1 3874731 90 + 14215200 ------AGCAGCCGGAGGAAGCAGAGCAUUCCAAGCGCAUUUAAUGCGCAUAAUGUCAAAUGCAACAAGGCAACGGA----AACAACAAACCUCAGUACA---------------- ------........((((..(((..(((((....((((((...))))))..)))))....))).....(....)(..----..)......))))......---------------- ( -22.30, z-score = -1.80, R) >droYak2.chr2L 18893961 90 + 22324452 ------GGAAGUCGGAGGAAGCAGAGCAUUCCAAGCGCAUUUAAUGCGCAUAAUGUCAAAUGCAACAAAGCAACGGA----AACAACAAACCUCAGUACA---------------- ------....(((.((((..(((..(((((....((((((...))))))..)))))....)))...........(..----..)......)))).).)).---------------- ( -22.10, z-score = -2.46, R) >droAna3.scaffold_13266 4729068 106 + 19884421 ------GAGAAGCGGAGGAAGUGGAGCAUUUCAAGCGCAUUUAAUGCGCAUAAUGUCAAAUGCAACAAAGCAACAAA----AACAACAGGCCACCACUCCUCCCAACCCACUCCCA ------.....(.((((((.((((.(((((....((((((...))))))..)))))....(((......))).....----............)))))))))))............ ( -29.70, z-score = -3.51, R) >dp4.chr3 12599257 75 - 19779522 ------AGAAGCUGGAGGAAAUGGAGCAUUUCAAGCGCAUUUAAUGCUCAUAAUGUCAAAUGCAACAAGAACCACAA----AACA------------------------------- ------....(((....((((((...)))))).)))((((((...((.......)).))))))..............----....------------------------------- ( -12.20, z-score = 0.09, R) >droPer1.super_2 1656533 75 - 9036312 ------AGAAGCUGGAGGAAAUGGAGCAUUUCAAGCGCAUUUAAUGCUCAUAAUGUCAAAUGCAACAAGAACCACAA----AACA------------------------------- ------....(((....((((((...)))))).)))((((((...((.......)).))))))..............----....------------------------------- ( -12.20, z-score = 0.09, R) >droVir3.scaffold_12875 16199715 97 + 20611582 GGCAGUUGCAAUGCCAGUUCGUGCAACAUUUCAAGCGCAUUUAAUGCGCAUAAUGUCAACUGCAAUGACUACAAGAGCUGUGGACACAAACCACAGU------------------- ((((.......))))((((..(((((((((....((((((...))))))..)))))....))))..))))......(((((((.......)))))))------------------- ( -31.40, z-score = -2.15, R) >droGri2.scaffold_15245 13022828 79 + 18325388 ---------------GGCAGUUGUAACAUUCGAAGCGCAUUUAAUGCGCCUAAUAUCAACUGU--UGGAGCCAGAAG----AAGCACAACUCACAGUUCA---------------- ---------------(..((((((....(((...((((((...))))))((((((.....)))--))).....))).----....))))))..)......---------------- ( -17.20, z-score = -0.52, R) >consensus ______AGAAGCCGGAGGAAGUGGAGCAUUCCAAGCGCAUUUAAUGCGCAUAAUGUCAAAUGCAACAAAGCAACGAA____AACAACAAACCACAGU_CA________________ .........................(((((....((((((...))))))..)))))............................................................ ( -7.10 = -7.29 + 0.19)


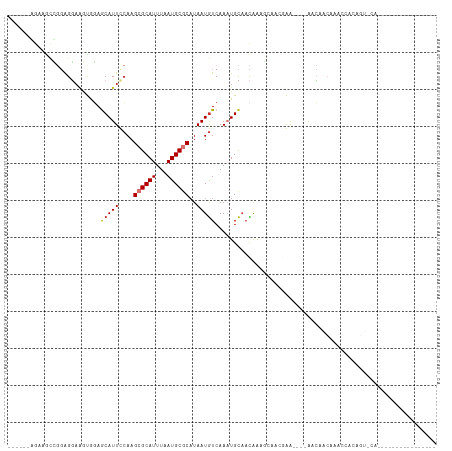
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:12:16 2011