| Sequence ID | dm3.chr2R |
|---|---|
| Location | 6,058,681 – 6,058,798 |
| Length | 117 |
| Max. P | 0.963440 |

| Location | 6,058,681 – 6,058,798 |
|---|---|
| Length | 117 |
| Sequences | 6 |
| Columns | 130 |
| Reading direction | forward |
| Mean pairwise identity | 69.93 |
| Shannon entropy | 0.51743 |
| G+C content | 0.39307 |
| Mean single sequence MFE | -26.27 |
| Consensus MFE | -12.49 |
| Energy contribution | -12.55 |
| Covariance contribution | 0.06 |
| Combinations/Pair | 1.29 |
| Mean z-score | -2.13 |
| Structure conservation index | 0.48 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.72 |
| SVM RNA-class probability | 0.963440 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
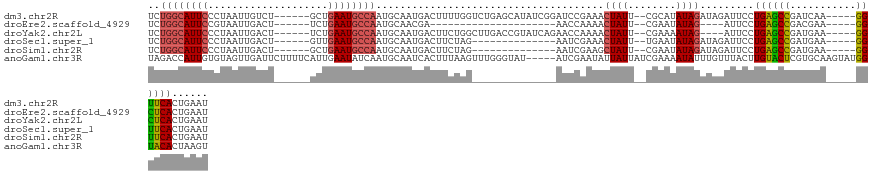
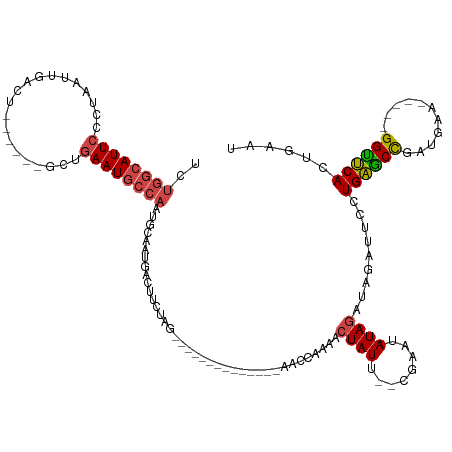
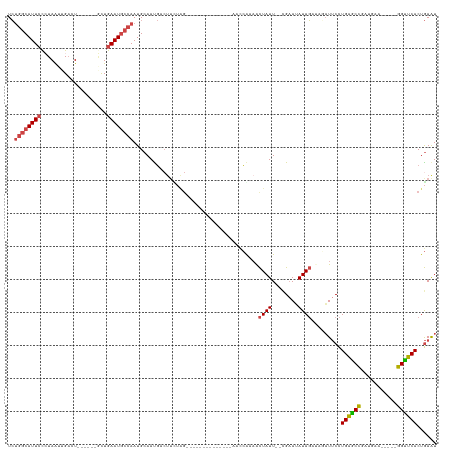
>dm3.chr2R 6058681 117 + 21146708 UCUGGCAUUCCCUAAUUGUCU------GCUGAAUGCCAAUGCAAUGACUUUUGGUCUGAGCAUAUCGGAUCCGAAACUAUU--CGCAUAUAGAUAGAUUCCUGAGCCGAUCAA-----GGUUCACUGAAU ..((((((((..((......)------)..))))))))((((((((..(((((((((((.....))))).)))))).))))--.))))........((((.((((((......-----))))))..)))) ( -33.70, z-score = -2.19, R) >droEre2.scaffold_4929 18330580 92 - 26641161 UCUGGCAUUCCGUAAUUGACU------UCUGAAUGCCAAUGCAACGA---------------------AACCAAAACUAUU--CGAAUAUAG----AUUCCUGAGCCGACGAA-----GGCUCACUGAAU ..((((((((.(.........------.).)))))))).........---------------------........((((.--.....))))----((((.((((((......-----))))))..)))) ( -23.20, z-score = -3.30, R) >droYak2.chr2L 18692906 113 + 22324452 UCUGGCAUUCCCUAAUUGACU------UCUGAAUGCCAAUGCAAUGACUUCUGGCUUGACCGUAUCAGAACCAAAACUAUU--CGAAAAUAG----AUUCCUGAGCCGAUGAA-----GGCUCACUGAAU ..((((((((...........------...))))))))..((...(....)..)).........((((........(((((--....)))))----.....((((((......-----)))))))))).. ( -26.24, z-score = -2.12, R) >droSec1.super_1 3674385 103 + 14215200 UCUGGCAUUCCCUAAUUGACU------GUUGAAUGCCAAUGCAAUGACUUCUAG--------------AAUCGAAACUAUU--UGAAUAUAGAUAGAUUCCUGAGCCGAUGAA-----GGUUCACUGAAU ..((((((((..((......)------)..))))))))..........(((..(--------------((((....((((.--.....))))...))))).((((((......-----))))))..))). ( -24.00, z-score = -2.29, R) >droSim1.chr2R 4690283 103 + 19596830 UCUGGCAUUCCCUAAUUGACU------GCUGAAUGCCAAUGCAAUGACUUCUAG--------------AAUCGAAGCUAUU--CGAAUAUAGAUAGAUUCCUGAGCCGAUGAA-----GGUUCACUGAAU ..((((((((..((......)------)..)))))))).........((((((.--------------..(((((....))--)))...)))).))((((.((((((......-----))))))..)))) ( -26.90, z-score = -2.34, R) >anoGam1.chr3R 42103460 125 + 53272125 UAGACCAUUGUGUAGUUGAUUCUUUUCAUUGAAUAUCAAUGCAAUCACUUUAAGUUUGGGUAU-----AUCGAAUAUUAUUAUCGAAAAUAUUUGUUUACUUGUACUCGUGCAAGUAUGGUACACUAAGU (((.(((...((.(((.((((.....(((((.....))))).))))))).))....)))((((-----..((((((((.........))))))))..((((((((....))))))))..)))).)))... ( -23.60, z-score = -0.52, R) >consensus UCUGGCAUUCCCUAAUUGACU______GCUGAAUGCCAAUGCAAUGACUUCUAG______________AACCAAAACUAUU__CGAAUAUAGAUAGAUUCCUGAGCCGAUGAA_____GGUUCACUGAAU ..((((((((....................))))))))...............................................................((((((...........))))))...... (-12.49 = -12.55 + 0.06)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:11:54 2011