
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 5,960,646 – 5,960,740 |
| Length | 94 |
| Max. P | 0.529303 |

| Location | 5,960,646 – 5,960,740 |
|---|---|
| Length | 94 |
| Sequences | 11 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 67.23 |
| Shannon entropy | 0.69773 |
| G+C content | 0.42819 |
| Mean single sequence MFE | -22.22 |
| Consensus MFE | -8.09 |
| Energy contribution | -8.16 |
| Covariance contribution | 0.08 |
| Combinations/Pair | 1.40 |
| Mean z-score | -1.14 |
| Structure conservation index | 0.36 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.07 |
| SVM RNA-class probability | 0.529303 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
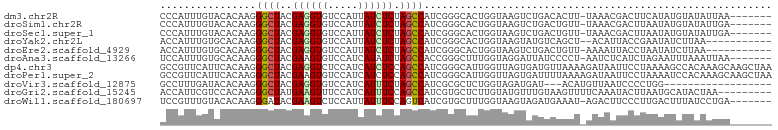
>dm3.chr2R 5960646 94 - 21146708 CCCAUUUGUACACAAGGGCUACGAGGUGUCCAUUAUCUCUAGCCAUCGGGCACUGGUAAGUCUGACACUU-UAAACGACUUCAUAUGUAUAUUAA------- ......((((((....(((((.((((((.....))))))))))).((((((........)))))).....-..............))))))....------- ( -24.10, z-score = -1.65, R) >droSim1.chr2R 4590720 94 - 19596830 CCCAUUUGUACACAAGGGCUACGAGGUGUCCAUUAUCUCUAGCCAUCGGGCACUGGUAAGUCUGACUGUU-UAAACGACUUAAUAUGUAUAUUGA------- ..((..((((((....(((((.((((((.....))))))))))).((((((........)))))).....-..............)))))).)).------- ( -24.60, z-score = -1.18, R) >droSec1.super_1 3573110 94 - 14215200 CCCAUUUGUACACAAGGGCUACGAGGUGUCCAUUAUCUCUAGCCAUCGGGCACUGGUAAGUCUGACUGUU-UAAACGACUUAAUAUGUAUAUUGA------- ..((..((((((....(((((.((((((.....))))))))))).((((((........)))))).....-..............)))))).)).------- ( -24.60, z-score = -1.18, R) >droYak2.chr2L 18593401 89 - 22324452 ACCAUUUGUGCACAAGGGCUACGAGGUGUCCAUUAUCUCUAGCCAUCGGGCACUGGUAAGUAUGUCAGCU--ACAUUACCGAAUAUCUUAA----------- .......((((.....(((((.((((((.....))))))))))).....))))(((((((((.(....))--)).))))))..........----------- ( -27.10, z-score = -2.32, R) >droEre2.scaffold_4929 18230449 90 + 26641161 ACCAUUUGUGCACAAGGGCUACGAGGUGUCCAUUAUCUCUAGCCAUCGGGCACUGGUAAGUCUGACUGUU-AAAAUUACCUAAUAUCUUAA----------- ((((...((((.....(((((.((((((.....))))))))))).....))))))))......((.((((-(........)))))))....----------- ( -24.20, z-score = -1.79, R) >droAna3.scaffold_13266 13816051 94 + 19884421 UCCAUUUGUGCACAAGGGCUACGAAGUGUCCAUCAUAUCUAGCCACCGGGCUUUGGUAGGAUUAUCCCCU-AAUCUCAUCUAGAAUUUAAAUUAA------- (((.((((....))))(((((.((.(((.....))).)))))))...))).(((((..((((((.....)-)))))...)))))...........------- ( -18.50, z-score = 0.14, R) >dp4.chr3 6173859 102 - 19779522 GCCGUUCAUUCACAAGGGCUACGAGGUCUCCAUCAUCUCCAGCCAUCGGGCAUUGGUUAGUGAUGUUAAAAGAUAAUUCCUAAAAGCCACAAAGCAAGCUAA ((.((......))...((((...((((((.((((((..((((((....)))..)))...)))))).....))).....)))...)))).....))....... ( -20.70, z-score = -0.02, R) >droPer1.super_2 6388733 102 - 9036312 GCCGUUCAUUCACAAGGGCUACGAAGUGUCCAUCAUCUCCAGCCAUCGGGCAUUGGUUAGUGAUUUUAAAAGAUAAUUCCUAAAAUCCACAAAGCAAGCUAA (((((......))...((((..((..((.....))..)).))))....))).(((.((.(((((((((...((....)).)))))).))).)).)))..... ( -19.00, z-score = -0.30, R) >droVir3.scaffold_12875 15042113 81 - 20611582 GCCUUUGAUACACAAGGGCUACGAGGUGUCCAUCAUUUCUAGCCAUCGCGCUCUGGUAGAUGAU---ACAUGUUAAUCCCCUGG------------------ (((((((.....)))))))...((.((((..(((((((...((((........)))))))))))---)))).))..........------------------ ( -20.20, z-score = -0.47, R) >droGri2.scaffold_15245 15067740 93 - 18325388 ACCAUUCGUCCACAAGGGCUAUGAAGUUUCCAUCAUUUCCAGCCAUCGUGCUCUUGUAUGUUUGUAAGUUUUCAAAUACUUAAUGCAUACUAA--------- .......(..(((...((((..(((((.......))))).))))...)))..)..((((((...(((((........)))))..))))))...--------- ( -20.20, z-score = -2.84, R) >droWil1.scaffold_180697 568805 94 + 4168966 UCCGUUUGUACACAAGGGAUACGAAGUCUCCAUUAUUUCCAGUCAUCGUGCUUUGGUAAGUAGAUGAAAU-AGACUUCCCUUGACUUUAUCCUGA------- ............(((((((.....(((((.............(((((.(((((....))))))))))...-))))))))))))............------- ( -21.19, z-score = -0.95, R) >consensus ACCAUUUGUACACAAGGGCUACGAGGUGUCCAUUAUCUCUAGCCAUCGGGCACUGGUAAGUCUGUCUGUU_UAAAAUACCUAAUAUCUAUAUUGA_______ ................((((..((((((.....)))))).)))).......................................................... ( -8.09 = -8.16 + 0.08)


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:11:41 2011