
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 5,835,432 – 5,835,582 |
| Length | 150 |
| Max. P | 0.880148 |

| Location | 5,835,432 – 5,835,582 |
|---|---|
| Length | 150 |
| Sequences | 7 |
| Columns | 157 |
| Reading direction | reverse |
| Mean pairwise identity | 60.38 |
| Shannon entropy | 0.77803 |
| G+C content | 0.41553 |
| Mean single sequence MFE | -40.62 |
| Consensus MFE | -11.94 |
| Energy contribution | -14.07 |
| Covariance contribution | 2.13 |
| Combinations/Pair | 1.65 |
| Mean z-score | -1.77 |
| Structure conservation index | 0.29 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.04 |
| SVM RNA-class probability | 0.880148 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 5835432 150 - 21146708 ------GUUUUUCUACAGCU-CCUAUAAAUGGUUUUUUCUUUCUAUUUUUUGCGCUUUUAGCGGCGUUUUAAAAGAGCAGUCGCGAAAGUGUUGCCUACUUUAUGGCGCGCACUUUUGGGUGUUAUUGGGGCCUAAUGGCUUCAGCUUGUGUGCAUU ------..............-.....((((((..........))))))..(((((....(((.((((..((((((.((((.(((....)))))))))..))))..))))(((((....)))))...(((((((....))))))))))...))))).. ( -43.00, z-score = -1.15, R) >droSim1.chr2R 4465684 146 - 19596830 ------GUUUUUCUACAGCU-CCUAUAAAUGGUUUU---UUUCUAUUUUCUGCGCUUUUAGCGGCGUUUUAAAAGAGCAGCCGCGAAAGUGUUGCCUACUUUAUGGCGCUCACUUUCGGGUGUUCUU-GUGCUUAAUGGCUGCAAGUUGGGUGCAUU ------...........((.-((((.((((((....---...))))))...((((((...(((((((((.....)))).)))))(((((((.((((........))))..))))))))))))).(((-(((((....))).))))).)))).))... ( -46.80, z-score = -2.45, R) >droSec1.super_1 3445450 145 - 14215200 ------GUUUUUCUACAGCU-CCUAUAAAUGGUUUU---UUUCUAUUUUCUGCGCUUUUAGCGGCG-UUUAAAAGAGCAGCCGCGAAAGUGUUGCCUACUUUAUGGCGCUCACUUUCGGGUGUUCUU-GUGCUUAAUGGCUGCAGGUUGGGUGCAUU ------...........((.-((((.((((((....---...))))))((((((((.(((..((((-.....(((((((.((.((((((((.((((........))))..)))))))))))))))))-.))))))).))).))))).)))).))... ( -49.10, z-score = -2.89, R) >droYak2.chr2L 18461686 136 - 22324452 ------GUUUUACUACAGCU-CCUAUAAACGGUUU----UUUCUA-UGUUC-CACUUUUGGUGGCGUUUUUAAAGAGCAGCCGCGAAAGUGUUGCCUACUUUAUGGCACUCACUUUUGGGUAUUCUU-GAGCUUAAUGGCUGCAGGCAUU------- ------...........(((-(....(((((....----......-....(-(((.....))))))))).....)))).(((.((((((((.((((........))))..)))))))))))...(((-(((((....)))).))))....------- ( -38.33, z-score = -1.57, R) >droEre2.scaffold_4929 18108379 137 + 26641161 ------GUUUUUCUACAGCU-CCUAUAAAUGUUUU----CUUCUAUUUUUC-CACUUUCGGUGGCGUUUUAAAAGAGCCACCGCGAAAGUGUUGCCUACUUUAUGGCGCUCACUUUGGGGUAUUCUU-GAGUUUAAUGGCUGCAUGCAUU------- ------.........(((((-....(((((.....----............-......(((((((...........)))))))(.((((((.((((........))))..)))))).).........-..)))))..)))))........------- ( -32.50, z-score = -0.79, R) >droAna3.scaffold_13266 15338287 150 - 19884421 GUUUUUGGAUUUUGGUAAUU-UUCAAGAGCAGGUUUGUGCAACAAAUCGUUGCA-UACUGUGGGGGGUAUAAAGGGAUUAUU-CGA---UAUUUUCUA-UUAAGACUGUGCUCUUGAGAGUAUCUUUAGGGUAUUGUGACUGCUCUUUCUUGCCAUU ............((((((((-((((((((((((((((((((((.....))))))-)...((((..(((((...(((....))-).)---))))..)))-)..))))).)))))))))))))......((((((.......))))))....))))).. ( -44.80, z-score = -3.30, R) >droGri2.scaffold_15245 7295671 123 + 18325388 --------------ACAAAUACAUAUAUAUAAAUCGAGACUUGUAGUUGCUGUGGCUCUCUGUUUGACUCUGUGAGUUUGCUGCAAAAGUGCUGCAUACCUUGUGGCGUUUGCUGUGUGUUUUGUAUUCGGCUCGAA-------------------- --------------.(.((((((.(((((((.....(((((..(((...(...(((.....))).)...)))..)))))...(((((...((..((.....))..)).))))))))))))..)))))).).......-------------------- ( -29.80, z-score = -0.24, R) >consensus ______GUUUUUCUACAGCU_CCUAUAAAUGGUUUU___UUUCUAUUUUUUGCGCUUUUAGCGGCGUUUUAAAAGAGCAGCCGCGAAAGUGUUGCCUACUUUAUGGCGCUCACUUUCGGGUAUUCUU_GGGCUUAAUGGCUGCAGGUUGU___CAUU ...............(((((...............................((((((((.(((((((((.....)))).)))))))))))))((((........)))).............................)))))............... (-11.94 = -14.07 + 2.13)
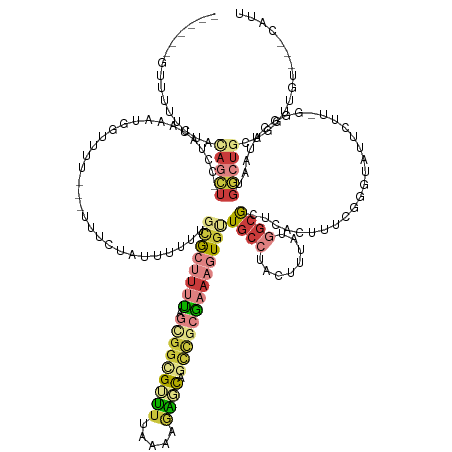


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:11:22 2011