| Sequence ID | dm3.chr2R |
|---|---|
| Location | 5,833,001 – 5,833,091 |
| Length | 90 |
| Max. P | 0.729184 |

| Location | 5,833,001 – 5,833,091 |
|---|---|
| Length | 90 |
| Sequences | 3 |
| Columns | 91 |
| Reading direction | reverse |
| Mean pairwise identity | 67.65 |
| Shannon entropy | 0.45313 |
| G+C content | 0.32963 |
| Mean single sequence MFE | -11.75 |
| Consensus MFE | -9.06 |
| Energy contribution | -7.73 |
| Covariance contribution | -1.33 |
| Combinations/Pair | 1.71 |
| Mean z-score | -0.84 |
| Structure conservation index | 0.77 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.52 |
| SVM RNA-class probability | 0.729184 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
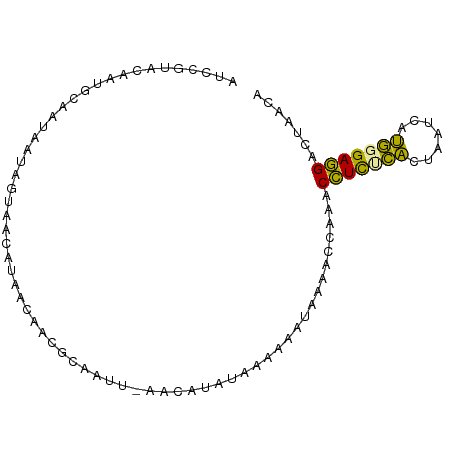
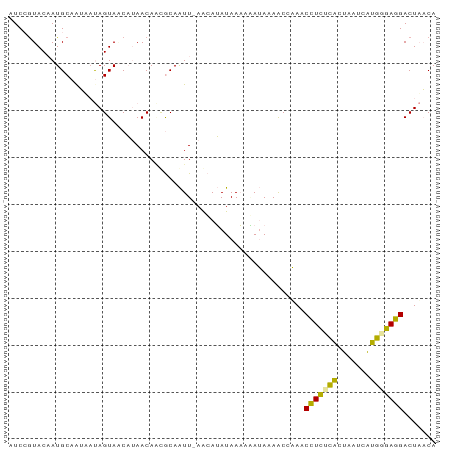
>dm3.chr2R 5833001 90 - 21146708 AUCAAUACAAUGCAAUAACAGUAACAUAACAGCGCAAUU-AACAUAUAAAAGAUAAAACCAAACCUCUCACUAAUCAUGUGAGGACUUACA ..........(((....................)))...-.........................((((((.......))))))....... ( -7.15, z-score = -0.17, R) >droEre2.scaffold_4820 1875259 90 + 10470090 AUCCGUAC-AUGCAGUACUAGUACCAUAACAUUGCAAUUAAACAUAUAAAUAAUAUAACCAAACUUCUCACUUAUCGUGGGAAGACUAACA ....((..-.((((((..((......))..))))))...........................((((((((.....))))))))....)). ( -15.40, z-score = -2.36, R) >droMoj3.scaffold_6473 7178164 90 + 16943266 AUCCGUACAAUGCAAGUAUAGUAGCUAAACAACACAGUU-AGAAAAUAAUAUAAAGAGCCUAACCUUCUGAUAUGCACAGGAGGACUAUCA ...............(.(((((.(((..........(((-(.....))))......)))....(((((((.......))))))))))))). ( -12.69, z-score = 0.01, R) >consensus AUCCGUACAAUGCAAUAAUAGUAACAUAACAACGCAAUU_AACAUAUAAAAAAUAAAACCAAACCUCUCACUAAUCAUGGGAGGACUAACA ...............................................................(((((((.......)))))))....... ( -9.06 = -7.73 + -1.33)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:11:21 2011