
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 2,446,804 – 2,446,949 |
| Length | 145 |
| Max. P | 0.733766 |

| Location | 2,446,804 – 2,446,949 |
|---|---|
| Length | 145 |
| Sequences | 6 |
| Columns | 152 |
| Reading direction | reverse |
| Mean pairwise identity | 81.38 |
| Shannon entropy | 0.34947 |
| G+C content | 0.38077 |
| Mean single sequence MFE | -38.20 |
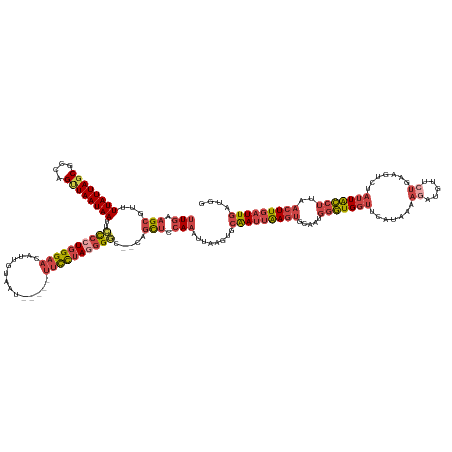
| Consensus MFE | -25.72 |
| Energy contribution | -27.95 |
| Covariance contribution | 2.23 |
| Combinations/Pair | 1.24 |
| Mean z-score | -1.64 |
| Structure conservation index | 0.67 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.54 |
| SVM RNA-class probability | 0.733766 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
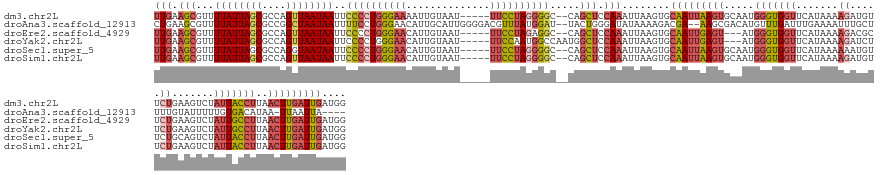
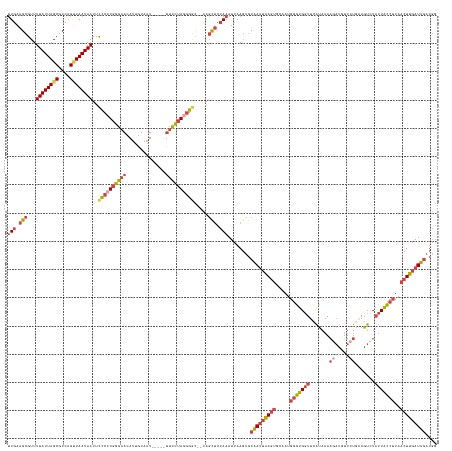
>dm3.chr2L 2446804 145 - 23011544 UUGAAGCGUUUUAUUAGCGCCAGUUAAUAAUUCCCCUGGGAAAAUUGUAAU-----UUCCUAGGGGC--CAGCUCCAAAUUAAGUGCAAUUAAGUGCAAUGGGUGGUUCAUAAAAGAUGUUCUGAAGUCUAUUACCUUAACUUGAUUGAUGG (((.(((...((((((((....))))))))..(((((((((((.......)-----)))))))))).--..))).)))........(((((((((.....(((((((.......((.....)).......)))))))..))))))))).... ( -44.24, z-score = -3.12, R) >droAna3.scaffold_12913 186250 143 + 441482 CUGAAGCGUUUUAUUAGCGCCGGCUAAUAAUUUUCCUGGGAACAUUGCAUUGGGGACGUUUAUGGAU--UACUGGGAUAUAAAAGACGA--AAGCGACAUGUUUGAUUUGAAAAUUUGCUUUUGUAUUUUUGUGACAUAA-UUAAUUA---- ..........((((((((....))))))))...(((..(...(((.....((....))...)))...--..)..)))((((((((((((--((((((...((((.......)))))))))))))).))))))))......-.......---- ( -31.70, z-score = -1.45, R) >droEre2.scaffold_4929 2490893 142 - 26641161 UUGAAGCGUUUUAUUAGCGCCAGUUAAUAAUUCCCCUGGGAACAUUGUAAU-----UUCCUAGAGGC--CAGCUCCAAAUUAAGUGCAAUUGAGU---AUGGGUGGUUCAUAAAAGACGCUCUGAAGUCUAUUGCCUUAACUUGAUUGAUGG (((.((((..((((((((....))))))))..)(((((((((.........-----))))))).)).--..))).)))........((((..(((---..(((..((.......((((........))))))..)))..)))..)))).... ( -38.51, z-score = -1.54, R) >droYak2.chr2L 2435872 144 - 22324452 UUGAAGCGUUUUAUUAGCGCCAGUUAAUAAUUCCCCUGGGAACAUUGUAAU-----UUCCAAUGGCCAAUGGCUCCAAAUUAAGUGCAAUUGAGU---AUGGGUGGUUCAUAAAAGAUCUUCUGAAGUCUAUUGCCUUAACUUGAUUGAUGG .....(((((.....)))))((.(((((..((((....))))(((((....-----...)))))(((...))).....))))).))((((..(((---..(((..((.......((.....)).......))..)))..)))..)))).... ( -34.04, z-score = -0.39, R) >droSec1.super_5 620388 145 - 5866729 UUGAAGCGUUUUAUUAGCGCCAGGUAAUAAUUCCCCUGGGAACAUUGUAAU-----UUCCUAGGGGC--CAGCUCCAAAUUAAGUGCAAUUAAGUGCAAUGGGUGGUUCAUAAAAAAUGUUCUGCAGUCUAUUACCUUAACUUGAUUGAUGG .....(((((.....))))).((((((((...((((((((((.........-----))))))))))(--(((((........)))(((......)))..)))(..(..(((.....)))..)..)....))))))))............... ( -38.00, z-score = -0.84, R) >droSim1.chr2L 2406151 145 - 22036055 UUGAAGCGUUUUAUUAGCGCCAGUUAAUAAUUCCCCUGGGAACAUUGUAAU-----UUCCUAGGGGC--CAGCUCCAAAUUAAGUGCAAUUAAGUGCAAUGGGUGGUUCAUAAAAGAUGUUCUGAAGUCUAUUACCUUAACUUGAUUGAUGG (((.(((...((((((((....))))))))..((((((((((.........-----)))))))))).--..))).)))........(((((((((.....(((((((.......((.....)).......)))))))..))))))))).... ( -42.74, z-score = -2.51, R) >consensus UUGAAGCGUUUUAUUAGCGCCAGUUAAUAAUUCCCCUGGGAACAUUGUAAU_____UUCCUAGGGGC__CAGCUCCAAAUUAAGUGCAAUUAAGUGCAAUGGGUGGUUCAUAAAAGAUGUUCUGAAGUCUAUUACCUUAACUUGAUUGAUGG (((.(((...((((((((....))))))))..((((((((((..............)))))))))).....))).)))........(((((((((.....(((((((.......((.....)).......)))))))..))))))))).... (-25.72 = -27.95 + 2.23)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:11:18 2011