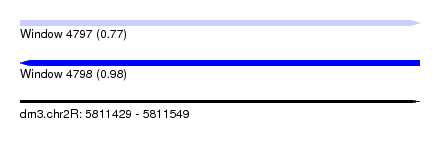

| Sequence ID | dm3.chr2R |
|---|---|
| Location | 5,811,429 – 5,811,549 |
| Length | 120 |
| Max. P | 0.975795 |

| Location | 5,811,429 – 5,811,549 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 93.49 |
| Shannon entropy | 0.11451 |
| G+C content | 0.48986 |
| Mean single sequence MFE | -37.29 |
| Consensus MFE | -33.78 |
| Energy contribution | -34.02 |
| Covariance contribution | 0.24 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.56 |
| Structure conservation index | 0.91 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.65 |
| SVM RNA-class probability | 0.774683 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
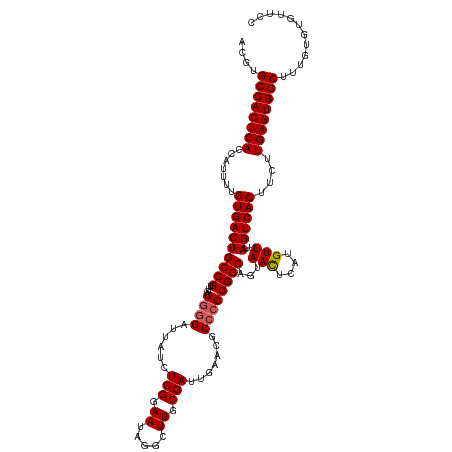
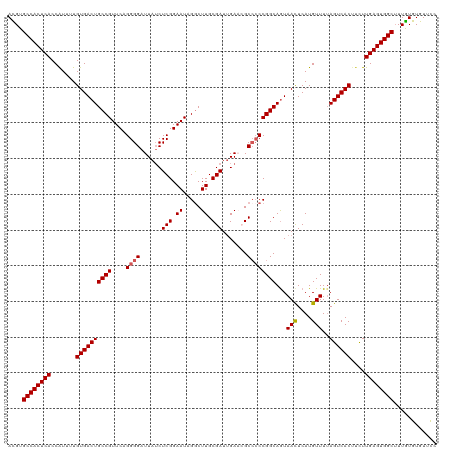
>dm3.chr2R 5811429 120 + 21146708 ACGUGCCACUCACCGUUUUGUGACUGCCGUUCAGGGGAUUAUCUCGGAAUAGGCUUGCGAUUGAACGCCCCCGGCAAUAACUCAUGUUUUAGUCACUUCUUGAGUGGCUUUGUGUGUUCC (((.((((((((.......((((((((((....((((......(((.((.....)).))).......))))))))...(((....)))..))))))....))))))))..)))....... ( -37.02, z-score = -1.44, R) >droEre2.scaffold_4929 18083887 118 - 26641161 AUGUGCCACUCACCAUUUUGUGACUGCCGUUCAGGGGAUUAGCUCGGAAUAGGCUUGCGAUGGA--GCAACCGGCAGUAAUUCAUGUUUUAGUCACCAUUUGAGUGGCAUUAUGUGUUCC ..((((((((((.......(((((((((((((.(((......))).)))).((.((((......--)))))))))).(((........))))))))....)))))))))).......... ( -37.80, z-score = -1.57, R) >droYak2.chr2L 18437395 118 + 22324452 AUGUGCCACUCACCAUUUUGUGACUGCCGUUCAGGGGAUUAUCUCGGAAUAGGCUUGCGAUUGA--GCCCCCGGCAGUAACUCAUGUUUAAGUCACUUCUUGAGUGGCUUUGUGUGUUUC ....((((((((.......(((((((((((((..(((.....))).)))).(((((......))--)))...)))...(((....)))..))))))....))))))))............ ( -37.60, z-score = -1.74, R) >droSec1.super_1 3421920 120 + 14215200 ACGUGCCACUCACCAUUUUGUGACUGCCGUUCAGGGGAUUAUCUCGGAAUAGGCUUGCGAUUGAACGCCCCCGGCAGUAACUCAUGUUUUAGUCACUUCUUGAGUGGCUUUGUGUGUUCC (((.((((((((.......((((((((((....((((......(((.((.....)).))).......))))))))...(((....)))..))))))....))))))))..)))....... ( -37.02, z-score = -1.32, R) >droSim1.chr2R 4442029 120 + 19596830 ACGUGCCACUCACCAUUUUGUGACUGCCGUUCAGGGGAUUAUCUCGGAAUAGACUUGCGAUUGAACGCCCCCGGCAGUAACUCAUGUUUUAGUCACUUCUUGAGUGGCUUUGUGUGUUCC (((.((((((((.......((((((((((....((((......(((.((.....)).))).......))))))))...(((....)))..))))))....))))))))..)))....... ( -37.02, z-score = -1.74, R) >consensus ACGUGCCACUCACCAUUUUGUGACUGCCGUUCAGGGGAUUAUCUCGGAAUAGGCUUGCGAUUGAACGCCCCCGGCAGUAACUCAUGUUUUAGUCACUUCUUGAGUGGCUUUGUGUGUUCC ....((((((((.......((((((((((....((((......(((.((.....)).))).......))))))))...(((....)))..))))))....))))))))............ (-33.78 = -34.02 + 0.24)

| Location | 5,811,429 – 5,811,549 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 93.49 |
| Shannon entropy | 0.11451 |
| G+C content | 0.48986 |
| Mean single sequence MFE | -37.14 |
| Consensus MFE | -33.26 |
| Energy contribution | -33.86 |
| Covariance contribution | 0.60 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.47 |
| Structure conservation index | 0.90 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.94 |
| SVM RNA-class probability | 0.975795 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
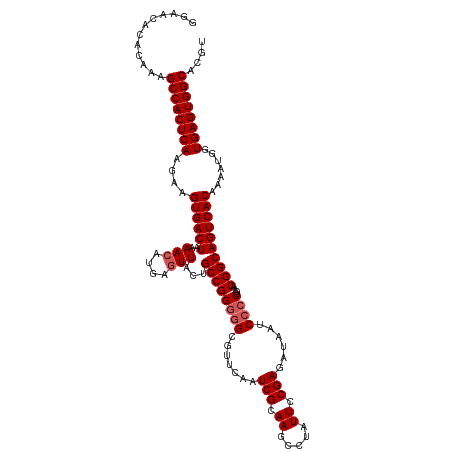
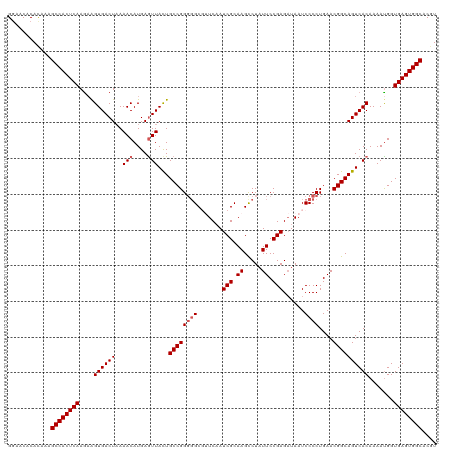
>dm3.chr2R 5811429 120 - 21146708 GGAACACACAAAGCCACUCAAGAAGUGACUAAAACAUGAGUUAUUGCCGGGGGCGUUCAAUCGCAAGCCUAUUCCGAGAUAAUCCCCUGAACGGCAGUCACAAAACGGUGAGUGGCACGU ............((((((((....((((((..(((....)))...((((((((.(((...(((.((.....)).)))...)))))))....)))))))))).......)))))))).... ( -37.10, z-score = -2.15, R) >droEre2.scaffold_4929 18083887 118 + 26641161 GGAACACAUAAUGCCACUCAAAUGGUGACUAAAACAUGAAUUACUGCCGGUUGC--UCCAUCGCAAGCCUAUUCCGAGCUAAUCCCCUGAACGGCAGUCACAAAAUGGUGAGUGGCACAU ...........(((((((((...(....).....(((.....(((((((..(((--......)))((((......).)))...........)))))))......))).)))))))))... ( -34.70, z-score = -2.35, R) >droYak2.chr2L 18437395 118 - 22324452 GAAACACACAAAGCCACUCAAGAAGUGACUUAAACAUGAGUUACUGCCGGGGGC--UCAAUCGCAAGCCUAUUCCGAGAUAAUCCCCUGAACGGCAGUCACAAAAUGGUGAGUGGCACAU ............((((((((...(((((((((....)))))))))((((((((.--....(((.((.....)).)))......))))....)))).............)))))))).... ( -38.30, z-score = -3.28, R) >droSec1.super_1 3421920 120 - 14215200 GGAACACACAAAGCCACUCAAGAAGUGACUAAAACAUGAGUUACUGCCGGGGGCGUUCAAUCGCAAGCCUAUUCCGAGAUAAUCCCCUGAACGGCAGUCACAAAAUGGUGAGUGGCACGU ............((((((((....((((((..(((....)))...((((((((.(((...(((.((.....)).)))...)))))))....)))))))))).......)))))))).... ( -37.10, z-score = -2.02, R) >droSim1.chr2R 4442029 120 - 19596830 GGAACACACAAAGCCACUCAAGAAGUGACUAAAACAUGAGUUACUGCCGGGGGCGUUCAAUCGCAAGUCUAUUCCGAGAUAAUCCCCUGAACGGCAGUCACAAAAUGGUGAGUGGCACGU ............((((((((....((((((..(((....)))...((((((((((......)))..((((......))))...)))).....))))))))).......)))))))).... ( -38.50, z-score = -2.53, R) >consensus GGAACACACAAAGCCACUCAAGAAGUGACUAAAACAUGAGUUACUGCCGGGGGCGUUCAAUCGCAAGCCUAUUCCGAGAUAAUCCCCUGAACGGCAGUCACAAAAUGGUGAGUGGCACGU ............((((((((....((((((..(((....)))...((((((((.......(((.((.....)).)))......))))....)))))))))).......)))))))).... (-33.26 = -33.86 + 0.60)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:11:17 2011