| Sequence ID | dm3.chr2R |
|---|---|
| Location | 5,677,040 – 5,677,132 |
| Length | 92 |
| Max. P | 0.947335 |

| Location | 5,677,040 – 5,677,132 |
|---|---|
| Length | 92 |
| Sequences | 11 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 75.89 |
| Shannon entropy | 0.49270 |
| G+C content | 0.36404 |
| Mean single sequence MFE | -17.79 |
| Consensus MFE | -14.04 |
| Energy contribution | -14.23 |
| Covariance contribution | 0.19 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.20 |
| Structure conservation index | 0.79 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.53 |
| SVM RNA-class probability | 0.947335 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 5677040 92 - 21146708 --------UUAGGAUCUGUGCUUUGAAUGUCAAAAUAUUU--UUGUGUUUGGCAUUUCAAGCGUUAAAUUUCUUCUGUCAACGCUCGUCCCCAGGAAAACUC --------...((((..((((((.((((((((((.(((..--..))))))))))))).))))(((..............)))))..))))............ ( -18.74, z-score = -1.21, R) >droSim1.chr2R 4308546 92 - 19596830 --------UUAGGAUCUGUGCUUUGAAUGUCAAAAUAUUU--UUGUGUUUGGCAUUUCAAGCGUUAAAUUUCUUCUGUCAACGCUCGUCCCCAGGAAAACUC --------...((((..((((((.((((((((((.(((..--..))))))))))))).))))(((..............)))))..))))............ ( -18.74, z-score = -1.21, R) >droSec1.super_1 3286034 92 - 14215200 --------UUAGGAUCUGUGCUUUGAAUGUCAAAAUAUUU--UUGUGUUUGGCAUUUCAAGCGUUAAAUUUCUUCUGUCAACGCUCGUCCCCAGGAAAACUC --------...((((..((((((.((((((((((.(((..--..))))))))))))).))))(((..............)))))..))))............ ( -18.74, z-score = -1.21, R) >droEre2.scaffold_4929 17952370 92 + 26641161 --------UUAGGAUCUGUGCUUUGAAUGUCAAAAUAUUU--UUGUGUUUGGCAUUUCAAGCGUUAAAUUUCUUCUGUCAACGCUCGUCCCUAGAAAAACUC --------...((((..((((((.((((((((((.(((..--..))))))))))))).))))(((..............)))))..))))............ ( -18.74, z-score = -1.85, R) >droYak2.chr2L 18301483 92 - 22324452 --------UUAGGAUCUGUGCUUUGAAUGUCAAAAUAUUU--UUGUGUUUGGCAUUUCAAGCGUUAAAUUUCUUCUGUCAACGCUCGUCCUGAGGAAAACUC --------(((((((..((((((.((((((((((.(((..--..))))))))))))).))))(((..............)))))..)))))))......... ( -22.44, z-score = -2.36, R) >droAna3.scaffold_13266 12947216 92 - 19884421 --------UUAGUAUCUGUGCUUUGAAUGUCAAAAUAUUU--GUGUGUUUGGCAUUUCAAGCGUUAAAUUUUUGCUGUCAACGCUCAACCCCAAAAAAACAA --------........(((((((.((((((((((.(((..--..))))))))))))).))))......((((((..((.........))..)))))).))). ( -16.60, z-score = -0.77, R) >dp4.chr3 14297821 90 - 19779522 -------UGUUGUAUCUGUGCUUUGAAUGUCAAAAUAUUU--GUGUGUUUGGCAUUUUAAGCGUUAAAUUUUGGCUGUCAACGUACCCUCAUUCGGCUC--- -------............((((.((((((((((.(((..--..))))))))))))).))))..........(((((................))))).--- ( -18.19, z-score = -1.01, R) >droPer1.super_4 6144352 90 + 7162766 -------UGUUGUAUCUGUGCUUUGAAUGUCAAAAUAUUU--GUGUGUUUGGCAUUUUAAGCGUUAAAUUUUGGCUGUCAACGUACCGUCAUUUGGCUC--- -------............((((.((((((((((.(((..--..))))))))))))).))))(((((((..(((...........)))..)))))))..--- ( -20.80, z-score = -1.77, R) >droWil1.scaffold_180701 3655374 84 - 3904529 --------UAUAUUUGUAUGCUUUGAAUUUCAAAAUAUUUGAGUGUGUUUGGCAUUUUAAGCGUUAAAUUCU--CAGUCAACAUUCCUUUUGCC-------- --------...(((((.((((((.((((.(((((((((....)))).))))).)))).)))))))))))...--....................-------- ( -11.60, z-score = 0.03, R) >droVir3.scaffold_12875 6003769 86 - 20611582 UGUGUGCUACUCGCUCUGUGCUCUGAAUGUCAAAAUAUUU--GUGUGUGUGGCAUUUUAAGCGUUAAAUUUCUGCUGUCAACAUUUCC-------------- .((((((.....)).....(((..((((((((..(((...--.)))...))))))))..)))..................))))....-------------- ( -14.90, z-score = -0.23, R) >droMoj3.scaffold_6496 2823069 86 + 26866924 UGCGCCUCACUCUCUCUGUGUUUUGAAUGUCAAAAUAUUU--GUGUGUUUGGCAUUUUAAGCGUUAAAUUUUUGCUGUCAACAUUUCC-------------- ...(((.(((.(.....((((((((.....))))))))..--).)))...)))......((((.........))))............-------------- ( -16.20, z-score = -1.55, R) >consensus ________UUAGGAUCUGUGCUUUGAAUGUCAAAAUAUUU__GUGUGUUUGGCAUUUCAAGCGUUAAAUUUCUGCUGUCAACGCUCCUCCCCAGGAAAAC__ ..................(((((.((((((((((.(((......))))))))))))).)))))....................................... (-14.04 = -14.23 + 0.19)
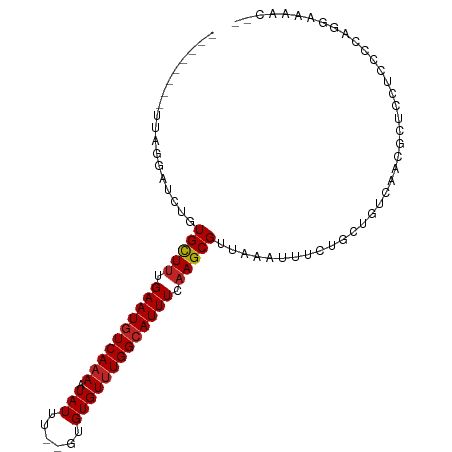


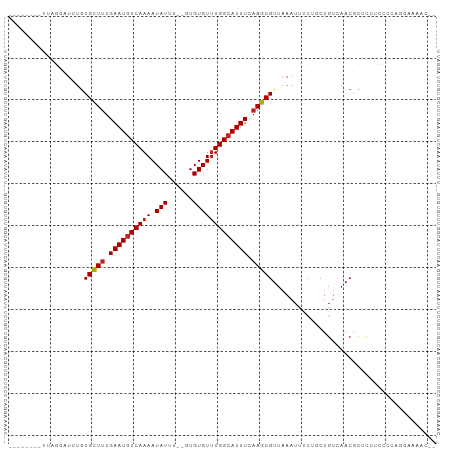
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:10:59 2011