| Sequence ID | dm3.chr2R |
|---|---|
| Location | 5,643,526 – 5,643,617 |
| Length | 91 |
| Max. P | 0.773656 |

| Location | 5,643,526 – 5,643,617 |
|---|---|
| Length | 91 |
| Sequences | 10 |
| Columns | 114 |
| Reading direction | forward |
| Mean pairwise identity | 71.84 |
| Shannon entropy | 0.54995 |
| G+C content | 0.59018 |
| Mean single sequence MFE | -38.25 |
| Consensus MFE | -16.02 |
| Energy contribution | -16.86 |
| Covariance contribution | 0.84 |
| Combinations/Pair | 1.27 |
| Mean z-score | -1.71 |
| Structure conservation index | 0.42 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.65 |
| SVM RNA-class probability | 0.773656 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
>dm3.chr2R 5643526 91 + 21146708 -----CUUUAAAUUAUGCCAGGUAUCGUCUGGCGUC---------------AGCCACGCCUCCUCCGCCCCCAAAUC---GGUGGGCGGGUGCUGUGUGCCGCUGGGAUUACCA -----.........((((((((.....))))))))(---------------((((((((..(.(((((((((.....---)).))))))).)..)))))..))))((....)). ( -38.30, z-score = -2.19, R) >droSim1.chr2R 4279851 91 + 19596830 -----CUUUAAAAUAUGCCAGGUAUCGUCUGGCGGC---------------AGCCACGCCUCCUCCGCCCCCAAAUC---GGUGGGCGGGUGCUGUGUGCCGCUGGGAUUACCA -----...............((((...((..(((((---------------(..(((((....(((((((((.....---)).))))))).)).)))))))))..))..)))). ( -42.00, z-score = -2.58, R) >droSec1.super_1 3252684 91 + 14215200 -----CUUUAAAAUAUGCCAGGUAUCGUCUGGCGGC---------------AGCCACGCCUCCUCCGCCCCCAAAUC---GGUGGGCGGGUGCUGUGUGCCGCUGGGAUUACCA -----...............((((...((..(((((---------------(..(((((....(((((((((.....---)).))))))).)).)))))))))..))..)))). ( -42.00, z-score = -2.58, R) >droYak2.chr2L 18269522 91 + 22324452 -----GUUUAAAAUAUGCCAGGUAUCGUCUGGCGGC---------------AACCACGCCUCCUCCGCCCCCAAAUC---GGUGGGCGGGUUCUGUGUGCCGCUGGGAUUACCA -----...............((((...((..(((((---------------(..((((.....(((((((((.....---)).)))))))...))))))))))..))..)))). ( -40.70, z-score = -2.96, R) >droEre2.scaffold_4929 17920276 91 - 26641161 -----GUUUAAAAUAUGCCAGGUAUCGUCUGGCGGC---------------AGCCACGCCUCCGCCGCCCCCAAAUC---GGUGGGCGGGUGCUGUGUGCCACUGGGAUUACCA -----...............((((...((..(.(((---------------(..(((((.(((((((((........---))).)))))).)).))))))).)..))..)))). ( -36.20, z-score = -0.46, R) >droAna3.scaffold_13266 12915902 91 + 19884421 -----UUUUUAAAUAUGCCAGAUAUCGUCUGGAGGC---------------AACAACUCCGCCAACGCCCCCAAAAC---GGUGGGCGGGUGCUGUGUGCCACUGGGAUUACCA -----............((((((...)))))).(((---------------(...((..((((..(((((((.....---)).)))))))))..)).))))...((.....)). ( -34.40, z-score = -1.95, R) >dp4.chr3 14267150 109 + 19779522 -----GUUUAAAAUAUGCCAGGUAUCGUCUGGAGGCGCCACACGCAGUGGCAGCAGCCGCACUCCAGCCCCCACAACAAUGGUGGGCGGCUGCUGUAUUCCACUGGGAUUACCG -----...............((((.((((....)))).....(.(((((((((((((((.....).(((((((......))).))))))))))))....)))))).)..)))). ( -43.40, z-score = -1.69, R) >droPer1.super_4 6114003 109 - 7162766 -----GUUUAAAAUAUGCCAGGUAUCGUCUGGAGGCGCCACACGCAGUGGGAGCAGCCGCACUCCAGCCCCCACAACAAUGGUGGGCGGCUGCUGUAUUCCACUGGGAUUACCG -----...............((((.((((....)))).....(.(((((((((((((.((......(((((((......))).)))).)).))))).)))))))).)..)))). ( -42.30, z-score = -1.32, R) >droWil1.scaffold_180701 3613439 111 + 3904529 CUUUAAAAAAAAAUCUGCCAGGUAUCGGCUGGUGGUGCCAAUGGCAAUGGCAACACCACCUCGAACACCUCAAAAAC---UGUAGGCGGCUGCUGUGUACCGCUGGGAUUACCG ...........(((((...((((.((((.(((((.(((((.......))))).))))).).)))..)))).......---....(((((.(((...)))))))).))))).... ( -31.80, z-score = -0.33, R) >droMoj3.scaffold_6496 2774872 76 - 26866924 ------------------------UGCUUUUGUGGCUC----------AGUUGCCGC-CCGGUGGCGCCCCUAAAAC---UGUGGGCGGCUGUUGUGUGCCGCUGGGAUUACCG ------------------------.......(((((..----------....)))))-(((((((((((((......---.).)))).((......))))))))))........ ( -31.40, z-score = -1.08, R) >consensus _____GUUUAAAAUAUGCCAGGUAUCGUCUGGCGGC_______________AGCCACGCCUCCUCCGCCCCCAAAAC___GGUGGGCGGGUGCUGUGUGCCGCUGGGAUUACCA .................((((.........((((......................)))).....(((((((........)).)))))..............))))........ (-16.02 = -16.86 + 0.84)


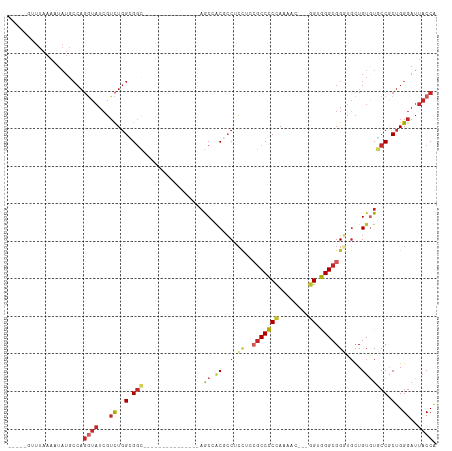
| Location | 5,643,526 – 5,643,617 |
|---|---|
| Length | 91 |
| Sequences | 10 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 71.84 |
| Shannon entropy | 0.54995 |
| G+C content | 0.59018 |
| Mean single sequence MFE | -36.86 |
| Consensus MFE | -14.80 |
| Energy contribution | -14.79 |
| Covariance contribution | -0.01 |
| Combinations/Pair | 1.47 |
| Mean z-score | -1.68 |
| Structure conservation index | 0.40 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.37 |
| SVM RNA-class probability | 0.664416 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
>dm3.chr2R 5643526 91 - 21146708 UGGUAAUCCCAGCGGCACACAGCACCCGCCCACC---GAUUUGGGGGCGGAGGAGGCGUGGCU---------------GACGCCAGACGAUACCUGGCAUAAUUUAAAG----- (((.....)))(((((.(((.((.(((((((.((---.....)))))))).)...))))))))---------------).)(((((.......)))))...........----- ( -36.80, z-score = -2.41, R) >droSim1.chr2R 4279851 91 - 19596830 UGGUAAUCCCAGCGGCACACAGCACCCGCCCACC---GAUUUGGGGGCGGAGGAGGCGUGGCU---------------GCCGCCAGACGAUACCUGGCAUAUUUUAAAG----- (((.....)))(((((.(((.((.(((((((.((---.....)))))))).)...))))))))---------------)).(((((.......)))))...........----- ( -39.70, z-score = -2.66, R) >droSec1.super_1 3252684 91 - 14215200 UGGUAAUCCCAGCGGCACACAGCACCCGCCCACC---GAUUUGGGGGCGGAGGAGGCGUGGCU---------------GCCGCCAGACGAUACCUGGCAUAUUUUAAAG----- (((.....)))(((((.(((.((.(((((((.((---.....)))))))).)...))))))))---------------)).(((((.......)))))...........----- ( -39.70, z-score = -2.66, R) >droYak2.chr2L 18269522 91 - 22324452 UGGUAAUCCCAGCGGCACACAGAACCCGCCCACC---GAUUUGGGGGCGGAGGAGGCGUGGUU---------------GCCGCCAGACGAUACCUGGCAUAUUUUAAAC----- .(((((((...(((.....).....((((((.((---.....)))))))).....))..))))---------------)))(((((.......)))))...........----- ( -34.30, z-score = -1.53, R) >droEre2.scaffold_4929 17920276 91 + 26641161 UGGUAAUCCCAGUGGCACACAGCACCCGCCCACC---GAUUUGGGGGCGGCGGAGGCGUGGCU---------------GCCGCCAGACGAUACCUGGCAUAUUUUAAAC----- (((.....)))(..((.(((.((.(((((((.((---.....)))))))).)...))))))).---------------.).(((((.......)))))...........----- ( -37.20, z-score = -1.49, R) >droAna3.scaffold_13266 12915902 91 - 19884421 UGGUAAUCCCAGUGGCACACAGCACCCGCCCACC---GUUUUGGGGGCGUUGGCGGAGUUGUU---------------GCCUCCAGACGAUAUCUGGCAUAUUUAAAAA----- (((.....)))..((((.(((((.(((((((.((---.....))))))).....)).))))))---------------))).(((((.....)))))............----- ( -30.10, z-score = -0.52, R) >dp4.chr3 14267150 109 - 19779522 CGGUAAUCCCAGUGGAAUACAGCAGCCGCCCACCAUUGUUGUGGGGGCUGGAGUGCGGCUGCUGCCACUGCGUGUGGCGCCUCCAGACGAUACCUGGCAUAUUUUAAAC----- .((((....((((((....((((((((((((.((((....)))))))).(.....)))))))))))))))(((.(((.....))).))).))))...............----- ( -46.20, z-score = -1.88, R) >droPer1.super_4 6114003 109 + 7162766 CGGUAAUCCCAGUGGAAUACAGCAGCCGCCCACCAUUGUUGUGGGGGCUGGAGUGCGGCUGCUCCCACUGCGUGUGGCGCCUCCAGACGAUACCUGGCAUAUUUUAAAC----- .((((....((((((.....(((((((((((.((((....)))))))).(.....)))))))).))))))(((.(((.....))).))).))))...............----- ( -43.00, z-score = -1.24, R) >droWil1.scaffold_180701 3613439 111 - 3904529 CGGUAAUCCCAGCGGUACACAGCAGCCGCCUACA---GUUUUUGAGGUGUUCGAGGUGGUGUUGCCAUUGCCAUUGGCACCACCAGCCGAUACCUGGCAGAUUUUUUUUUAAAG .((.....)).(((((........)))))....(---((((.(.((((((.((.(.(((((.(((((.......))))).))))).))))))))).).)))))........... ( -36.90, z-score = -2.12, R) >droMoj3.scaffold_6496 2774872 76 + 26866924 CGGUAAUCCCAGCGGCACACAACAGCCGCCCACA---GUUUUAGGGGCGCCACCGG-GCGGCAACU----------GAGCCACAAAAGCA------------------------ .((.....)).(((((........))))).....---(((((.(.((((((....)-))(....).----------..))).).))))).------------------------ ( -24.70, z-score = -0.32, R) >consensus UGGUAAUCCCAGCGGCACACAGCACCCGCCCACC___GAUUUGGGGGCGGAGGAGGCGUGGCU_______________GCCGCCAGACGAUACCUGGCAUAUUUUAAAC_____ ........((((..............(((((.((........))))))).....((((..((................)))))).........))))................. (-14.80 = -14.79 + -0.01)



Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:10:56 2011