
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 5,643,039 – 5,643,131 |
| Length | 92 |
| Max. P | 0.721394 |

| Location | 5,643,039 – 5,643,131 |
|---|---|
| Length | 92 |
| Sequences | 8 |
| Columns | 97 |
| Reading direction | reverse |
| Mean pairwise identity | 76.95 |
| Shannon entropy | 0.43307 |
| G+C content | 0.34357 |
| Mean single sequence MFE | -18.83 |
| Consensus MFE | -6.80 |
| Energy contribution | -7.30 |
| Covariance contribution | 0.50 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.51 |
| Structure conservation index | 0.36 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.50 |
| SVM RNA-class probability | 0.721394 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
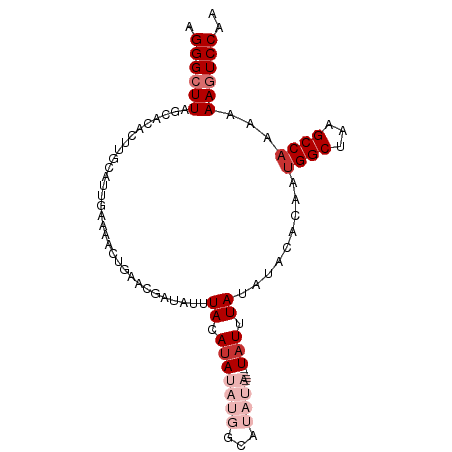
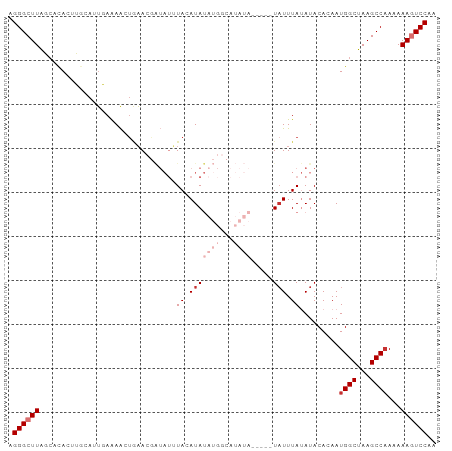
>dm3.chr2R 5643039 92 - 21146708 AGGGCUUUGCACACUUGCAGUGAAAAGUGAACGAUAUUUACAUAUAUGGCAUAUA-----UAUUUAUAUACACAAUGGCAAAGCCAAAAAAGUCCAA ..((((((((.(((((........)))))...........(((...((..(((((-----....))))).))..)))))))))))............ ( -21.90, z-score = -2.37, R) >droSim1.chr2R 4279362 92 - 19596830 AGGGCUUAGCACACUUGCAGUGAAAAGUGAACGAUAUUUACAUAUAUGGCAUAUA-----UAUUUAUAUACACAAUGGCUAAGCCAAAAAAGUCCAA ..((((((((.(((((........)))))...........(((...((..(((((-----....))))).))..)))))))))))............ ( -22.30, z-score = -2.58, R) >droSec1.super_1 3252180 92 - 14215200 AGGGCUUAGCACACUUGCAGUGAAAAGUGAACGAUAUUUACAUAUAUGGCAUAUA-----UAUUUAUAUACACAAUGGCUAAGCCAAAAAAGUCCAA ..((((((((.(((((........)))))...........(((...((..(((((-----....))))).))..)))))))))))............ ( -22.30, z-score = -2.58, R) >droYak2.chr2L 18269054 96 - 22324452 UGGGCUUAGCACACUUGCAUUGAAAACUGAACGAUAUUUACAUAUAUGGCAUAUAUAUAGUAUUUAUAUACACAAUGGCUAAGCC-AAAAAGUCCAA ..((((((((.((.(((((.(....).))....(((..(((.((((((.....)))))))))..))).....)))))))))))))-........... ( -20.70, z-score = -1.94, R) >droEre2.scaffold_4929 17919800 92 + 26641161 AGGGCUUAGCACACUUGCAUUGAAAACUGAACGAUAUUUACAUAUAUGGCAUAUA-----UUUUUAUAUACACAAUGGCUAAGCCAAAAAAGUCCAA ..((((((((.(....)(((((.........(.((((......)))).).(((((-----....)))))...)))))))))))))............ ( -18.70, z-score = -1.99, R) >droAna3.scaffold_13266 12915339 82 - 19884421 AGGGCUUAAAAUAAACACGCACAAAAAACUACUGUAUAUACAUAU---------------UAUUUAUAUACACAAUGGCUAAGCCAAAAAAGUCCGA .((((((.........................((((((((.....---------------....))))))))...((((...))))...)))))).. ( -16.10, z-score = -3.28, R) >dp4.chr3 14266588 74 - 19779522 GGGGCUUACCACACAGCCAUUCAGAGCUAAA-------UACAUA----------------UAUUUAAAUACACAAUGGCUAAGCCAAAAAACUCCAG ..(((((.......(((((((...(..((((-------((....----------------))))))..)....))))))))))))............ ( -14.31, z-score = -2.68, R) >droPer1.super_4 6113441 74 + 7162766 GGGGCUUACCACACAGCCAUUCAGAGCUAAA-------UACAUA----------------UAUUUAAAUACACAAUGGCUAAGCCAAAAAACUCCAG ..(((((.......(((((((...(..((((-------((....----------------))))))..)....))))))))))))............ ( -14.31, z-score = -2.68, R) >consensus AGGGCUUAGCACACUUGCAUUGAAAACUGAACGAUAUUUACAUAUAUGGCAUAUA_____UAUUUAUAUACACAAUGGCUAAGCCAAAAAAGUCCAA .((((((.....................................................(((....))).....((((...))))...)))))).. ( -6.80 = -7.30 + 0.50)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:10:55 2011