
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 5,582,403 – 5,582,500 |
| Length | 97 |
| Max. P | 0.613271 |

| Location | 5,582,403 – 5,582,500 |
|---|---|
| Length | 97 |
| Sequences | 9 |
| Columns | 99 |
| Reading direction | reverse |
| Mean pairwise identity | 79.89 |
| Shannon entropy | 0.41086 |
| G+C content | 0.51065 |
| Mean single sequence MFE | -29.22 |
| Consensus MFE | -17.51 |
| Energy contribution | -17.27 |
| Covariance contribution | -0.24 |
| Combinations/Pair | 1.19 |
| Mean z-score | -1.44 |
| Structure conservation index | 0.60 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.25 |
| SVM RNA-class probability | 0.613271 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 5582403 97 - 21146708 CGCGCAGCUGUAGGCGUAAAUUAAAAUGCAUUUUACGCAACUGUG-CGACAAGAUCCGGGCUUUAGAUCCUGCAGCUGUUG-CCCCUCGUACUGCAUUU .(((((((((((((((((((...........)))))))....((.-((........)).))........))))))))).))-)................ ( -29.30, z-score = -0.93, R) >droSim1.chr2R 4224292 97 - 19596830 CGCGCAGCUGUAGGCGUAAAUUAAAAUGCAUUUUACGCAACUGUG-CGACGAGAUCCGGGCUUCAGAUCCUGCAGCUGUUG-CCCCUCGUACUGCAUUC (((((((......(((((((...........)))))))..)))))-))(((((....((((..(((.........)))..)-))))))))......... ( -32.00, z-score = -1.53, R) >droSec1.super_1 3200180 97 - 14215200 CGCGCAGCUGUAGGCGUAAAUUAAAAUGCAUUUUACGCAACUGUG-CGACGAGAUACGGGCUUCAGAUCCUGCAGCUGUUG-UCCCUCGUACUGCAUUC (((((((......(((((((...........)))))))..)))))-))(((((..(((((((.(((...))).))))..))-)..)))))......... ( -30.40, z-score = -1.12, R) >droYak2.chr2L 18214786 92 - 22324452 CGCGCAGCUGUAGGCGUAAAUUAAAAUGCAUUUUACGCAACUGUG-CGACGAGAUCCGGGCUUCGGAUCCUGCAGCUGUUG-CCCCUCGCAUUU----- .(((((((((((((((((((...........))))))).......-......((((((.....))))))))))))))).))-)...........----- ( -34.50, z-score = -2.25, R) >droEre2.scaffold_4929 17869544 92 + 26641161 CGCGCAGCUGUAGGCGUAAAUUAAAAUGCAUUUUACGCAACUGUG-CGACGAGAUCCGGGCUGCAGAUCCUGCAGCUGUUG-CCCCUCGUAGUU----- (((((((......(((((((...........)))))))..)))))-))(((((...((((((((((...))))))))..))-...)))))....----- ( -34.30, z-score = -2.08, R) >droAna3.scaffold_13266 12867410 93 - 19884421 CGCGCAGCUGUAGCUGUAAAUUAAAAUGCAUUUUACGCAACUGUU-CACCAGGAUCCGGGCUAUAGAUCCUGCAGCUGUUGCCCCCUUGCAUUU----- ...(((((....)))))......(((((((......(((((.(((-...(((((((.........))))))).))).))))).....)))))))----- ( -29.80, z-score = -2.19, R) >dp4.chr3 14215845 98 - 19779522 CGCGCAGCUGUAGCUGUAAAUUAAAAUGCAUUUUACGCAACUGUUCCGUUGAUCCUCGAUGCUCGGAUCCUGCAGCUGUUG-CCGCUCGUGCUACCCCC .(((((((.(((((((((.(((.....(((((.....((((......))))......)))))...)))..)))))))...)-).))).))))....... ( -27.10, z-score = -1.61, R) >droPer1.super_4 6062644 98 + 7162766 CGCGCAGCUGUAGCUGUAAAUUAAAAUGCAUUUUACGCAACUGUUCCGUUGAUCCUCGAUGCUCGGAUCCUGCAGCUGUUG-CCGCUCGUGCUACCCCC .(((((((.(((((((((.(((.....(((((.....((((......))))......)))))...)))..)))))))...)-).))).))))....... ( -27.10, z-score = -1.61, R) >droWil1.scaffold_180701 3538221 87 - 3904529 --CGCAGCUGUAACUGUAAAUUAAAAUGCAUUUUACGCAACUGU--UCCUAAGCCAAAGAGGUCUCGUCCUGCAGCUGUUG-CGGCACAUUC------- --....((((((((............(((.......))).((((--......(((.....)))........))))..))))-))))......------- ( -18.44, z-score = 0.34, R) >consensus CGCGCAGCUGUAGGCGUAAAUUAAAAUGCAUUUUACGCAACUGUG_CGACGAGAUCCGGGCUUCAGAUCCUGCAGCUGUUG_CCCCUCGUACUGC____ ...((((((((((.((((((...........))))))...(((...((........)).....)))...)))))))))).................... (-17.51 = -17.27 + -0.24)

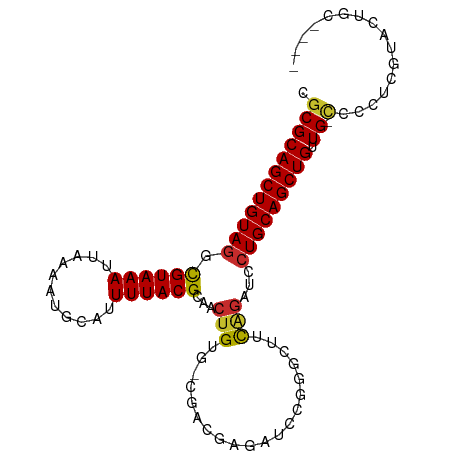

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:10:47 2011