
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 5,116,850 – 5,117,002 |
| Length | 152 |
| Max. P | 0.999667 |

| Location | 5,116,850 – 5,116,970 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 72.65 |
| Shannon entropy | 0.48651 |
| G+C content | 0.38929 |
| Mean single sequence MFE | -27.52 |
| Consensus MFE | -15.17 |
| Energy contribution | -16.69 |
| Covariance contribution | 1.52 |
| Combinations/Pair | 1.29 |
| Mean z-score | -1.47 |
| Structure conservation index | 0.55 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.71 |
| SVM RNA-class probability | 0.795443 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
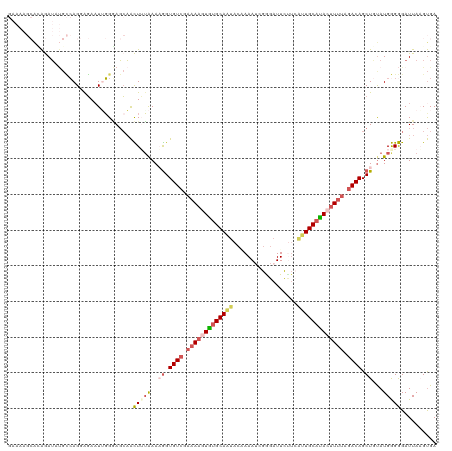
>dm3.chr2R 5116850 120 + 21146708 GAAAAGCACAGAUAUGCCAGGAGAAUUGGGAGCAACUUUAAAAGGUGUCGAAAAGUGGGCAAUACAAAAACGAGAUUCUUCAUUGCAUACUUUUAGACAGCUGUAUGGGGGAUUAAGUGA ......(((.(...(.((((.....)))).).)..(((((..((.((((.(((((((.(((((.(......).((....))))))).))))))).)))).))...)))))......))). ( -25.70, z-score = -0.91, R) >droSim1.chr2R 3781223 120 + 19596830 GAAAAGCACAGAUAUGCCAGGAGAAUGGGGAGCAACUUUAAAAGGUGUCGAAAAGUGUGCAAUACGAAAUCGAGAUUCUUCAUUGCAUACUUUUAGACAGCUGUAUGGGGGAUUAAGUGA ......(((.....(.(((......))).).....(((((..((.((((.(((((((((((((.((....)).((....))))))))))))))).)))).))...)))))......))). ( -32.80, z-score = -2.79, R) >droSec1.super_1 2748419 120 + 14215200 GAAAAGCAUAGAUAUGCCAGGAGAUUUGGGAGCAACUAUAAAAGGUGUCGAAAAGUGUGCAAUACCAAAUCGGGAUUCUUCAUUGCACACUUUUAGACAGCUGUAUGGGGGAUUAAGUGA .....((((....))))......((((((...(..(((((..((.((((.(((((((((((((.((.....))((....))))))))))))))).)))).)).)))))..).)))))).. ( -36.30, z-score = -3.45, R) >droYak2.chr2L 17771735 105 + 22324452 ---------GAAGACCACAGUGUAAUAGGAACAUGUUGUGAAUGGUGUCGAAAAGUAUGCAUU--UAAAUUGGGCUUC----GUGCAUAAUUGGAAACAGCUCUAUGAGAGAUUAUGCGA ---------((((.((.((((.(((((......)))))(((((((((.(.....)))).))))--)).))))))))))----.((((((((((....)).(((.....))))))))))). ( -23.60, z-score = -0.68, R) >droEre2.scaffold_4929 9099690 112 - 26641161 -------GAAAAUUAGUAAGCAGAAUUUGUAGCAAUUUGAAAUAGUGUCGAAAAGUAUGCAUU-CAAAAUCGGGAUUUCUGGUUGCAUAAGUUUAGACAGCUCCAUGAGAGAUUAUGUGA -------...........(((.(((((......))))).......((((.(((..((((((..-((((((....)))).))..))))))..))).))))))).(((((....)))))... ( -19.20, z-score = 0.49, R) >consensus GAAAAGCACAGAUAUGCCAGGAGAAUUGGGAGCAACUUUAAAAGGUGUCGAAAAGUGUGCAAUACAAAAUCGGGAUUCUUCAUUGCAUACUUUUAGACAGCUGUAUGGGGGAUUAAGUGA ...................................(((((..((.((((.(((((((((((((..................))))))))))))).)))).))...))))).......... (-15.17 = -16.69 + 1.52)
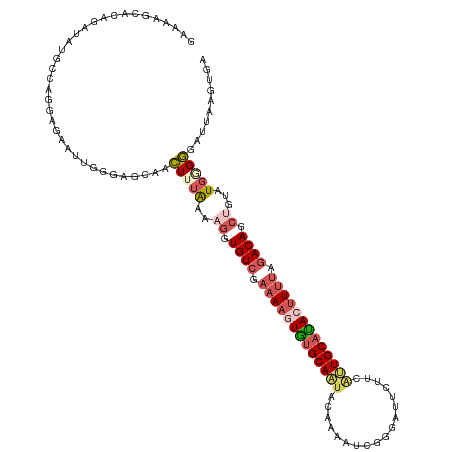


| Location | 5,116,850 – 5,116,970 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 72.65 |
| Shannon entropy | 0.48651 |
| G+C content | 0.38929 |
| Mean single sequence MFE | -21.20 |
| Consensus MFE | -14.31 |
| Energy contribution | -16.72 |
| Covariance contribution | 2.41 |
| Combinations/Pair | 1.17 |
| Mean z-score | -2.36 |
| Structure conservation index | 0.68 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.95 |
| SVM RNA-class probability | 0.996582 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
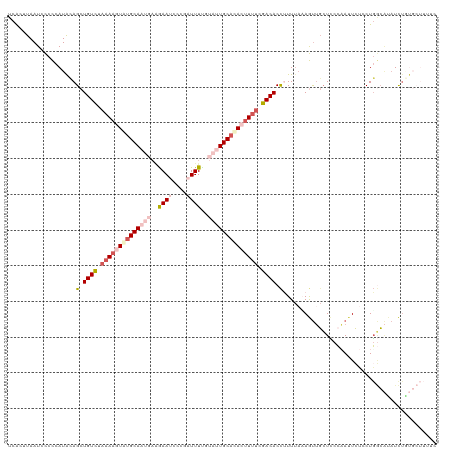
>dm3.chr2R 5116850 120 - 21146708 UCACUUAAUCCCCCAUACAGCUGUCUAAAAGUAUGCAAUGAAGAAUCUCGUUUUUGUAUUGCCCACUUUUCGACACCUUUUAAAGUUGCUCCCAAUUCUCCUGGCAUAUCUGUGCUUUUC .................(((.((((.((((((..((((((.((((......)))).))))))..)))))).))))........(((((....)))))...)))((((....))))..... ( -21.00, z-score = -2.17, R) >droSim1.chr2R 3781223 120 - 19596830 UCACUUAAUCCCCCAUACAGCUGUCUAAAAGUAUGCAAUGAAGAAUCUCGAUUUCGUAUUGCACACUUUUCGACACCUUUUAAAGUUGCUCCCCAUUCUCCUGGCAUAUCUGUGCUUUUC ...............(((((.((((.((((((.(((((((..((((....))))..))))))).)))))).))))...........((((............))))...)))))...... ( -24.20, z-score = -3.08, R) >droSec1.super_1 2748419 120 - 14215200 UCACUUAAUCCCCCAUACAGCUGUCUAAAAGUGUGCAAUGAAGAAUCCCGAUUUGGUAUUGCACACUUUUCGACACCUUUUAUAGUUGCUCCCAAAUCUCCUGGCAUAUCUAUGCUUUUC ..................(((((((.((((((((((((((..((((....))))..)))))))))))))).))))......((((.((((............))))...))))))).... ( -31.80, z-score = -4.87, R) >droYak2.chr2L 17771735 105 - 22324452 UCGCAUAAUCUCUCAUAGAGCUGUUUCCAAUUAUGCAC----GAAGCCCAAUUUA--AAUGCAUACUUUUCGACACCAUUCACAACAUGUUCCUAUUACACUGUGGUCUUC--------- ..((((((((((.....))).........)))))))..----((((.((......--..............((......))...((((((.......))).))))).))))--------- ( -13.40, z-score = -0.18, R) >droEre2.scaffold_4929 9099690 112 + 26641161 UCACAUAAUCUCUCAUGGAGCUGUCUAAACUUAUGCAACCAGAAAUCCCGAUUUUG-AAUGCAUACUUUUCGACACUAUUUCAAAUUGCUACAAAUUCUGCUUACUAAUUUUC------- .................((((((((.(((..((((((..(((((.......)))))-..))))))..))).))))........((((......))))..))))..........------- ( -15.60, z-score = -1.50, R) >consensus UCACUUAAUCCCCCAUACAGCUGUCUAAAAGUAUGCAAUGAAGAAUCCCGAUUUUGUAUUGCACACUUUUCGACACCUUUUAAAGUUGCUCCCAAUUCUCCUGGCAUAUCUGUGCUUUUC ...................(.((((.((((((((((((((..((((....))))..)))))))))))))).)))).)........................................... (-14.31 = -16.72 + 2.41)


| Location | 5,116,890 – 5,117,002 |
|---|---|
| Length | 112 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 72.27 |
| Shannon entropy | 0.47232 |
| G+C content | 0.35560 |
| Mean single sequence MFE | -29.58 |
| Consensus MFE | -17.90 |
| Energy contribution | -19.35 |
| Covariance contribution | 1.45 |
| Combinations/Pair | 1.25 |
| Mean z-score | -2.82 |
| Structure conservation index | 0.61 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 3.25 |
| SVM RNA-class probability | 0.998061 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
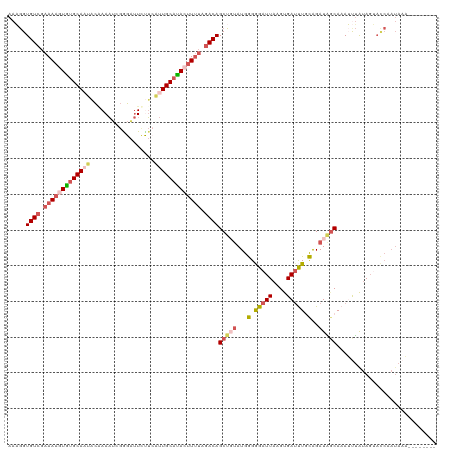
>dm3.chr2R 5116890 112 + 21146708 AAAGGUGUCGAAAAGUGGGCAAUACAAAAACGAGAUUCUUCAUUGCAUACUUUUAGACAGCUGUAUGGGGGAUUAAGUGAUUUGCAGAUGGCAAAUUUAUAGUACCCAUAAA-------- ..((.((((.(((((((.(((((.(......).((....))))))).))))))).)))).)).((((((..((((...(((((((.....)))))))..)))).))))))..-------- ( -33.20, z-score = -3.90, R) >droSim1.chr2R 3781263 112 + 19596830 AAAGGUGUCGAAAAGUGUGCAAUACGAAAUCGAGAUUCUUCAUUGCAUACUUUUAGACAGCUGUAUGGGGGAUUAAGUGAUUUGCAGAUAGCAAAUUUAUAGUACCAAUAAA-------- ..((.((((.(((((((((((((.((....)).((....))))))))))))))).)))).))...(((...((((...(((((((.....)))))))..)))).))).....-------- ( -35.10, z-score = -4.80, R) >droSec1.super_1 2748459 113 + 14215200 AAAGGUGUCGAAAAGUGUGCAAUACCAAAUCGGGAUUCUUCAUUGCACACUUUUAGACAGCUGUAUGGGGGAUUAAGUGAUUUGCAGAUAGCAAAUUUAUAGUACCAAUAAAA------- ..((.((((.(((((((((((((.((.....))((....))))))))))))))).)))).))...(((...((((...(((((((.....)))))))..)))).)))......------- ( -37.30, z-score = -5.04, R) >droYak2.chr2L 17771766 114 + 22324452 AAUGGUGUCGAAAAGUAUGCA-UUUAAAUU--GGGCUUCG---UGCAUAAUUGGAAACAGCUCUAUGAGAGAUUAUGCGAUCUUUGGAUAGCAAAACUGUAGAAUCCAUAAUUUUCAACA .....(((.(((((.((((.(-(((.....--..(((...---((((((((((....)).(((.....)))))))))))(((....)))))).........)))).)))).))))).))) ( -21.89, z-score = 0.25, R) >droEre2.scaffold_4929 9099723 100 - 26641161 AAUAGUGUCGAAAAGUAUGCA-UUCAAAAUCGGGAUUUCUGGUUGCAUAAGUUUAGACAGCUCCAUGAGAGAUUAUGUGAACUUUGGAUUCCAAAACAAAA------------------- .....((((.(((..((((((-..((((((....)))).))..))))))..))).))))..(((((((....))))).))..(((((...)))))......------------------- ( -20.40, z-score = -0.63, R) >consensus AAAGGUGUCGAAAAGUGUGCAAUACAAAAUCGGGAUUCUUCAUUGCAUACUUUUAGACAGCUGUAUGGGGGAUUAAGUGAUUUGCAGAUAGCAAAUUUAUAGUACCAAUAAA________ .....((((.(((((((((((((..................))))))))))))).))))(((((...(.(((((....))))).)..)))))............................ (-17.90 = -19.35 + 1.45)


| Location | 5,116,890 – 5,117,002 |
|---|---|
| Length | 112 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 72.27 |
| Shannon entropy | 0.47232 |
| G+C content | 0.35560 |
| Mean single sequence MFE | -25.93 |
| Consensus MFE | -14.63 |
| Energy contribution | -17.48 |
| Covariance contribution | 2.85 |
| Combinations/Pair | 1.21 |
| Mean z-score | -3.76 |
| Structure conservation index | 0.56 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 4.16 |
| SVM RNA-class probability | 0.999667 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
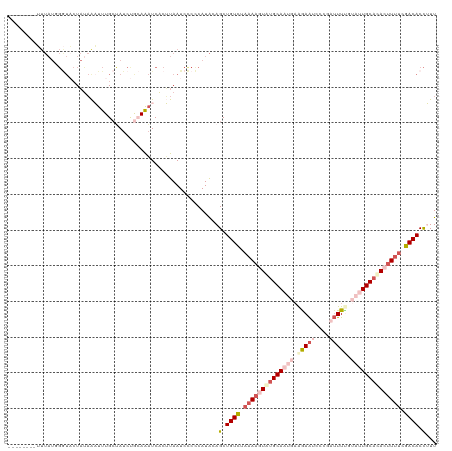
>dm3.chr2R 5116890 112 - 21146708 --------UUUAUGGGUACUAUAAAUUUGCCAUCUGCAAAUCACUUAAUCCCCCAUACAGCUGUCUAAAAGUAUGCAAUGAAGAAUCUCGUUUUUGUAUUGCCCACUUUUCGACACCUUU --------..((((((........((((((.....))))))..........)))))).((.((((.((((((..((((((.((((......)))).))))))..)))))).)))).)).. ( -26.77, z-score = -4.18, R) >droSim1.chr2R 3781263 112 - 19596830 --------UUUAUUGGUACUAUAAAUUUGCUAUCUGCAAAUCACUUAAUCCCCCAUACAGCUGUCUAAAAGUAUGCAAUGAAGAAUCUCGAUUUCGUAUUGCACACUUUUCGACACCUUU --------......(((.......((((((.....)))))).....................(((.((((((.(((((((..((((....))))..))))))).)))))).))))))... ( -26.20, z-score = -4.56, R) >droSec1.super_1 2748459 113 - 14215200 -------UUUUAUUGGUACUAUAAAUUUGCUAUCUGCAAAUCACUUAAUCCCCCAUACAGCUGUCUAAAAGUGUGCAAUGAAGAAUCCCGAUUUGGUAUUGCACACUUUUCGACACCUUU -------.......(((.......((((((.....)))))).....................(((.((((((((((((((..((((....))))..)))))))))))))).))))))... ( -32.80, z-score = -5.70, R) >droYak2.chr2L 17771766 114 - 22324452 UGUUGAAAAUUAUGGAUUCUACAGUUUUGCUAUCCAAAGAUCGCAUAAUCUCUCAUAGAGCUGUUUCCAAUUAUGCA---CGAAGCCC--AAUUUAAA-UGCAUACUUUUCGACACCAUU (((((((((...((((....((((((((((.(((....))).)))............))))))).))))..((((((---.(((....--..)))...-)))))).)))))))))..... ( -24.80, z-score = -2.68, R) >droEre2.scaffold_4929 9099723 100 + 26641161 -------------------UUUUGUUUUGGAAUCCAAAGUUCACAUAAUCUCUCAUGGAGCUGUCUAAACUUAUGCAACCAGAAAUCCCGAUUUUGAA-UGCAUACUUUUCGACACUAUU -------------------......(((((...)))))((((.(((........)))))))((((.(((..((((((..(((((.......)))))..-))))))..))).))))..... ( -19.10, z-score = -1.69, R) >consensus ________UUUAUGGGUACUAUAAAUUUGCUAUCUGCAAAUCACUUAAUCCCCCAUACAGCUGUCUAAAAGUAUGCAAUGAAGAAUCCCGAUUUUGUAUUGCACACUUUUCGACACCUUU ...........................................................(.((((.((((((((((((((.(((((....))))).)))))))))))))).)))).)... (-14.63 = -17.48 + 2.85)


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:09:51 2011