
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 4,950,007 – 4,950,183 |
| Length | 176 |
| Max. P | 0.850093 |

| Location | 4,950,007 – 4,950,183 |
|---|---|
| Length | 176 |
| Sequences | 6 |
| Columns | 186 |
| Reading direction | forward |
| Mean pairwise identity | 75.43 |
| Shannon entropy | 0.44905 |
| G+C content | 0.43873 |
| Mean single sequence MFE | -45.98 |
| Consensus MFE | -29.42 |
| Energy contribution | -31.43 |
| Covariance contribution | 2.01 |
| Combinations/Pair | 1.35 |
| Mean z-score | -1.44 |
| Structure conservation index | 0.64 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.91 |
| SVM RNA-class probability | 0.850093 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 4950007 176 + 21146708 --AUAUAUAUUGUAUGAUAUAAAUAGCUCGUGCAGUUGC------GAUUAAAAAUAAGAUCAGGAGUUCCUC--GAACGACUGAAGUGUUGCCACCGUAAAAAGAGCGGAAUGCUUCCUCAUUUUCGUCAUUUUCCUGACGGCAGCACGGCAAGAAAUCCGCUUUAGGAGCUCCUGGCAUCCACGG --...((((((....)))))).......((((....(((------((((........))))((((((((((.--((.((..((..((((((((.((((.......)))).................((((......))))))))))))..)).(....))).)).)))))))))).)))..)))). ( -52.30, z-score = -2.00, R) >droSim1.chr2R 3620242 171 + 19596830 -------AUAUAUAUGAUAUAAAUAGCUUGUGCAGUUGC------GAUUAAAAAUAGGAUCGGGAGUUCCUC--GAAUGACUGAAGUGUUGCCACCGUAAAAAGAGCGGAAUGCUUCCUCAUUUUCGUCAUUUUCCUGACGGCAGCACGGCAAGAAAUCCGCUUUAGGAGCUCCUGGCAUCCACGG -------.......((((...(((.((....)).)))..------.))))......((((((((((((((((--(.((..(((..((((((((.((((.......)))).................((((......))))))))))))..).))..)).))....)))))))))))..)))).... ( -51.10, z-score = -1.47, R) >droSec1.super_1 2588498 171 + 14215200 -------AUAUAUAUGAUAUAAAUAGCUUGUGCAGUCGC------GAUUAAAAAUAAGAUCGGGAGUUCCUC--GAAUGACUGAAGUGUUGCCACCGUAAAAAGAGCGGAAUGUUUCCUCAUUUUCGUCAUUUUCCUGACGGCAGCACGGCAAGAAAUCCGCUUUAGGAGCUCCUGGCAUCCACGG -------.....................((((..(((.(------((((........)))))((((((((((--(.((..(((..((((((((.((((.......)))).................((((......))))))))))))..).))..)).))....))))))))).)))...)))). ( -48.20, z-score = -1.12, R) >droYak2.chr2L 17611490 175 + 22324452 ---------UUAAUUGUUAUAAAUAGCUUAUGCAGCCACAGCCACGAUCAAAAAUAAGAUCAGGAGUUCCUC--GAAUGACUGAAGUGUUGUCACCGUAAAAAGAACGGAAUGCUUCCUCAUUUUCGUCAUAUUCCUGACGGCAGCACGGCAAGAAAUCCGCUUUAGGAGCUCCUGGCAUCCACAG ---------.....(((........((....)).......(((..((((........))))(((((((((((--(.((..(((..((((((((.((((.......)))).................((((......))))))))))))..).))..)).))....)))))))))))))....))). ( -49.30, z-score = -2.47, R) >droEre2.scaffold_4929 8941959 169 - 26641161 ---------AUAAUUGUUAUAAAUAGCGUAAGCAGCCGC------GAUUAAAAAUAAGAUCAAGAGUUCCUC--GAAUGACUGAAGUGUGGCCACCGUAAAAAGAGCGGAAUGCUUCCUCAUUUUCGUCAUAUUCCUGACGGCACCACGGCAAGAAAUCCGCUUUAGGAGCUCCUGGCAUCCACGG ---------................((....)).(((((------((((........))))..(((...)))--.............)))))..((((....((((((((...(((((......((((((......))))))......)).)))...)))))))).(((((.....)).))))))) ( -46.20, z-score = -1.29, R) >droAna3.scaffold_13266 16338952 173 - 19884421 AUAAAUACUGAAUUAUUUAUAAACAGCUUUAUAGACUUC-----CAGGCACUUCCCAAUUCAGGAAAUUCUCCGAAACCCCCGAAGUAUUGACACCUUAUAUAGGACAGAGUUCUUCCUAUUUUUUGUAAUUUUCGUCCUUCCAGCUG---AAACCAUCCACAGUCCAGACGGCAAGUCGC----- (((((((......)))))))...(((((.....(((...-----(((..(((((....(((.(((.....))))))......))))).)))(((......((((((.((....))))))))....))).......))).....)))))---.................(((.....)))..----- ( -28.80, z-score = -0.29, R) >consensus ________UAUAUAUGAUAUAAAUAGCUUGUGCAGCCGC______GAUUAAAAAUAAGAUCAGGAGUUCCUC__GAAUGACUGAAGUGUUGCCACCGUAAAAAGAGCGGAAUGCUUCCUCAUUUUCGUCAUUUUCCUGACGGCAGCACGGCAAGAAAUCCGCUUUAGGAGCUCCUGGCAUCCACGG .........................((....)).(((........((((........))))((((((((((..............((((((((.((((.......)))).................((((......))))))))))))((........)).....)))))))))))))........ (-29.42 = -31.43 + 2.01)

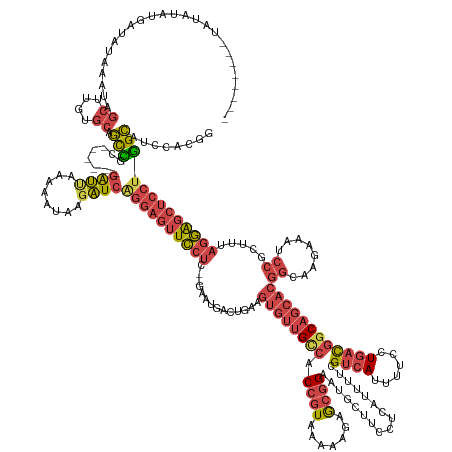

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:09:33 2011