
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 4,568,455 – 4,568,559 |
| Length | 104 |
| Max. P | 0.520956 |

| Location | 4,568,455 – 4,568,559 |
|---|---|
| Length | 104 |
| Sequences | 14 |
| Columns | 108 |
| Reading direction | forward |
| Mean pairwise identity | 80.47 |
| Shannon entropy | 0.44305 |
| G+C content | 0.46347 |
| Mean single sequence MFE | -28.21 |
| Consensus MFE | -14.82 |
| Energy contribution | -13.96 |
| Covariance contribution | -0.85 |
| Combinations/Pair | 1.54 |
| Mean z-score | -1.45 |
| Structure conservation index | 0.53 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.05 |
| SVM RNA-class probability | 0.520956 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 4568455 104 + 21146708 CAAGACCAACGAGGAUUUCUUUGAGGAGGAUGGACAGAGAUGGUUUAAAACCGGCGACAUUGGCGAAAUACAAGCUGAUGGCGUACUUAAGAUUAUUGGUAAGU---- ....((((((((((....)))))..(((............((((.....))))(((.(((..((.........))..))).))).))).......)))))....---- ( -22.70, z-score = -1.33, R) >droVir3.scaffold_12875 11413868 104 + 20611582 UAAAACCAAUGAGGACUUUUUUGAUGAAGAUGGACGCCGUUGGUUCAAAACCGGCGAUAUUGGAGAAGUGCAAUCCGAUGGCGUACUUAAGAUAAUUGGUAUGU---- ....((((((.....((((......))))....(((((((((((.....))))))...((((((.........)))))))))))..........))))))....---- ( -27.50, z-score = -2.01, R) >droGri2.scaffold_15245 5015067 104 - 18325388 CAAGACCAAUGAGGACUUUUUCGAGGAAGACGGACGCCGUUGGUUCAAAACCGGCGACAUUGGAGAAGUGCAAACCGAUGGCGUACUUAAGAUAAUUGGUGCGU---- ....((((((.....(((((....)))))....((((((((((((((...((((.....)))).....))..))))))))))))..........))))))....---- ( -26.90, z-score = -0.57, R) >droMoj3.scaffold_6496 11354706 104 - 26866924 AAAGACCAAUGAGGACUUUUUCGAAGAGGAUGGUCGCCGUUGGUUCAAAACCGGCGACAUUGGCGAAGUGCACUCAGAUGGCGUACUUAAGAUUAUUGGUGAGU---- ....((((((((.......((((..((.....(((((((((.......)).))))))).))..))))((((.((.....)).))))......))))))))....---- ( -28.70, z-score = -0.82, R) >droWil1.scaffold_180700 1644150 104 + 6630534 CAAGACCAAUGAAGAUUUCUUCGAGGAAGAUGGUCGUCGGUGGUUCAAAACUGGUGACAUUGGUGAAGUGCAAAGUGAUGGUGUACUUAAAAUUAUUGGUAGGU---- ....((((((((.....(((((...)))))..(((..((((........))))..))).......(((((((.(....)..)))))))....))))))))....---- ( -27.20, z-score = -2.16, R) >droPer1.super_4 5551350 108 - 7162766 CAAGACCAACGAGGAUUUCUUCGAGGAGGAUGGCAAACGCUGGUUCAAAACCGGCGACAUUGGCGAGGUGCAAACCGAUGGCGUACUUAAGAUAAUUGGCAAGUCUCG .........(((((....)))))..((((.((.(((.(((((((.....))))))).((((((...........))))))...............))).))..)))). ( -30.00, z-score = -0.73, R) >dp4.chr3 13709790 108 + 19779522 CAAGACCAACGAGGAUUUCUUCGAGGAGGAUGGCAAACGCUGGUUCAAAACCGGCGACAUUGGCGAGGUGCAAACCGAUGGCGUACUUAAGAUAAUUGGCAAGUCUCG .........(((((....)))))..((((.((.(((.(((((((.....))))))).((((((...........))))))...............))).))..)))). ( -30.00, z-score = -0.73, R) >droAna3.scaffold_13266 18439741 106 - 19884421 CAAGACCAACGAGGACUUCUUCGAGGAGGAUGGCAGACGGUGGUUCAAAACUGGAGACAUUGGAGAAAUCCAAGCCGAUGGCGUACUUAAGAUUAUUGGUAAGAAU-- ....((((((((((....)))))..(((.(((.((..((((..........((....))(((((....))))))))).)).))).))).......)))))......-- ( -31.50, z-score = -2.84, R) >droEre2.scaffold_4929 8568545 104 - 26641161 CAAGACCAACGAAGAUUUCUUCGAGGAAGAUGGGCAAAGAUGGUUCAAAACCGGCGACAUUGGCGAAAUACAAGCUGAUGGCGUACUUAAGAUUAUUGGUAAGU---- ....((((((((((....)))))...(((.((.((...(.((((.....)))).)..(((..((.........))..))))).))))).......)))))....---- ( -25.30, z-score = -1.72, R) >droYak2.chr2L 17228886 104 + 22324452 CAAGACCAAUGAAGAUUUCUUUGAGGAGGAUGGACAAAGAUGGUUCAAAACCGGCGACAUUGGCGAAAUACACGCUGAUGGCGUACUUAAGAUUAUUGGUAAGU---- ....((((((((.....((((.(((..((.(((((.......)))))...)).(((.(((..(((.......)))..))).))).)))))))))))))))....---- ( -27.90, z-score = -2.94, R) >droSec1.super_1 2212934 104 + 14215200 CAAGACCAACGAGGAUUUCUUUGAGGAGGAUGGGUUGAGAUGGUUUAAAACCGGCGACAUUGGCGAAAUUCAAGCUGAUGGCGUACUUAAGAUUAUUGGUAAGU---- ....(((..(((((....))))).....(((((.(((((.((((.....))))(((.(((..((.........))..))).))).)))))..))))))))....---- ( -24.40, z-score = -1.18, R) >droSim1.chr2R 3230396 104 + 19596830 CAAGACCAACGAGGAUUUCUUUGAGGAGGAUGGGCAGAGAUGGUUUAAAACCGGCGACAUUGGCGAAAUUCAAGCUGAUGGCGUACUUAAGAUUAUUGGUAAGU---- ....((((((((((....)))))..(((............((((.....))))(((.(((..((.........))..))).))).))).......)))))....---- ( -22.70, z-score = -0.59, R) >anoGam1.chr2R 42544905 108 + 62725911 CAAAACGCAGGAAGAUUUCUUCGAGGAGGACGGCCAGCGAUGGUUCCGUACCGGUGACGUGGGCGAAAUCCAUCCCGAUGGUGCGCUUAAGAUUAUCGGUAAGUGCAG ......(((.((((....)))).....(((..((((....))))))).(((((((((..((((((....((((....))))..))))))...)))))))))..))).. ( -36.40, z-score = -1.18, R) >triCas2.ChLG5 11610377 104 + 18847211 UAAAACAGACGAAGAUUUCACGGAUGUCGACGGCAGGAGGUGGUUCAGGACCGGCGAUAUCGGGGAAAUCCACCCCGAUGGUGUCGUUAAAAUCAUCGGUAAGU---- .............((((((.(.(((((((.((((..(((....)))..).))).))))))).).))))))....((((((((.........)))))))).....---- ( -33.80, z-score = -1.47, R) >consensus CAAGACCAACGAGGAUUUCUUCGAGGAGGAUGGCCAACGAUGGUUCAAAACCGGCGACAUUGGCGAAAUGCAAGCCGAUGGCGUACUUAAGAUUAUUGGUAAGU____ ....((((((((((....)))))...(((...........((((.....))))(((.((((((...........)))))).))).))).......)))))........ (-14.82 = -13.96 + -0.85)

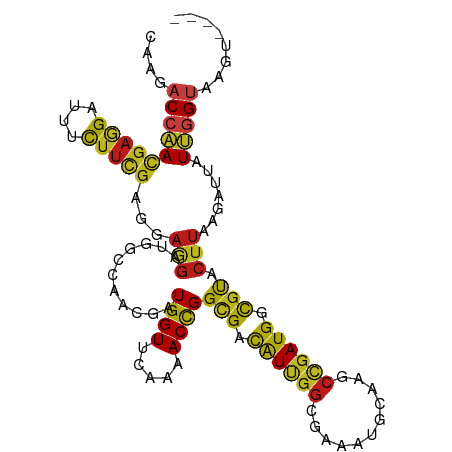

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:08:35 2011