
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 4,502,947 – 4,503,003 |
| Length | 56 |
| Max. P | 0.882597 |

| Location | 4,502,947 – 4,503,003 |
|---|---|
| Length | 56 |
| Sequences | 4 |
| Columns | 65 |
| Reading direction | forward |
| Mean pairwise identity | 62.81 |
| Shannon entropy | 0.56950 |
| G+C content | 0.32887 |
| Mean single sequence MFE | -10.03 |
| Consensus MFE | -7.05 |
| Energy contribution | -4.92 |
| Covariance contribution | -2.13 |
| Combinations/Pair | 1.90 |
| Mean z-score | -0.76 |
| Structure conservation index | 0.70 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.06 |
| SVM RNA-class probability | 0.882597 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
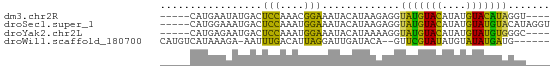
>dm3.chr2R 4502947 56 + 21146708 -----CAUGAAUAUGACUCCAAACGGAAAUACAUAAGAGGUAUGUACAUAUGUACAUAGGU---- -----......((((..(((....)))....)))).....(((((((....)))))))...---- ( -9.90, z-score = -1.00, R) >droSec1.super_1 2146541 60 + 14215200 -----CAUGGAAAUGACUCCAAAUGGAAAUACAUAAGAGGUAUGUACAUAUGUAUGUACAUAGGU -----(((....))).(((...(((......)))..))).(((((((((....)))))))))... ( -13.90, z-score = -1.96, R) >droYak2.chr2L 17159106 56 + 22324452 -----CAUGAGAAUGACUCCAAAUGGAAAUACAUAAAAGGUAUGUACAUAUGUAUGUGGGC---- -----((((...(((..(((....)))..((((((.....)))))))))...)))).....---- ( -8.00, z-score = -0.09, R) >droWil1.scaffold_180700 1556382 56 + 6630534 CAUGUCAUAAAGA-AAUUUGACAUUAGGAUUGAUACA--GUUCGUAUAUGUAUAUGAUG------ ...((((((((..-..))..((((..((((((...))--))))....)))).)))))).------ ( -8.30, z-score = -0.01, R) >consensus _____CAUGAAAAUGACUCCAAAUGGAAAUACAUAAAAGGUAUGUACAUAUGUAUGUAGGU____ .................(((....))).............(((((((....)))))))....... ( -7.05 = -4.92 + -2.13)


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:08:28 2011