
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 4,423,017 – 4,423,128 |
| Length | 111 |
| Max. P | 0.521541 |

| Location | 4,423,017 – 4,423,128 |
|---|---|
| Length | 111 |
| Sequences | 7 |
| Columns | 116 |
| Reading direction | forward |
| Mean pairwise identity | 65.69 |
| Shannon entropy | 0.69086 |
| G+C content | 0.49654 |
| Mean single sequence MFE | -32.10 |
| Consensus MFE | -6.28 |
| Energy contribution | -6.61 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.56 |
| Mean z-score | -2.18 |
| Structure conservation index | 0.20 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.05 |
| SVM RNA-class probability | 0.521541 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 4423017 111 + 21146708 CACAAACUUUCGCCUUCAUUGACGCCUGCAAACCGGUGAGCCCUUCGGAAGCGC-CCACUCAAACAGCCUGGUUAUAAAAUCA-GGCCAAUUAGUGCACG---UGAGAAAGGGGAC .........((.((((..((.(((..((((....((((..((....))...)))-)..........((((((((....)))))-))).......))))))---).)).)))).)). ( -30.20, z-score = -0.40, R) >droSim1.chr2R 3059888 111 + 19596830 CACAAACUUUCGCCUUCAUUGACGCCUGCAAACCGGUGAGCCCUUCGGAAGCGC-CCACUCAAACAGCCUGGUUAUAAAAUCA-GGCCAAUUAGUGCACG---UGAGAAAGGCGAC .........(((((((..((.(((..((((....((((..((....))...)))-)..........((((((((....)))))-))).......))))))---).)).))))))). ( -34.30, z-score = -2.13, R) >droSec1.super_1 2064944 111 + 14215200 CACAAACUUUCGCCUUCAUUGACGCCUGCAAACCGGUGAGCCCUUCGGAAGCGG-CCACUCAAACAGCCUGGUUAUAAAAUCA-GGCCAAUUAGUGCACG---UGAGAAAGGCGAC .........(((((((..((.(((..((((....((((.(((...(....).))-)))))......((((((((....)))))-))).......))))))---).)).))))))). ( -36.90, z-score = -2.48, R) >droYak2.chr2L 17078151 95 + 22324452 CACAAACUUUCGCCUUCAUCGACGCCUGCAAUUCCGUG-----------------CAACUCAAACAGCCUGGUUAUAAAAUCA-GGCCAAUUAGUGCACG---UGAGAAAGGCGAC .........(((((((..(((..(....).....((((-----------------((.((......((((((((....)))))-))).....))))))))---)))..))))))). ( -31.50, z-score = -4.01, R) >droEre2.scaffold_4929 8422293 111 - 26641161 CCCGAACUUUCGCCUUCAUCGCCGCCUGCAAAACUGUGAGCCUCUCGGAAGCUC-CUACUCAAAAAGCCUGGUUAUAAAAUCA-GGCCAAUUAGUGCACG---UGAGAUAGGCGAC .........((((((..(((..(((.((((.......((((.((...)).))))-...........((((((((....)))))-))).......)))).)---)).))))))))). ( -34.10, z-score = -2.52, R) >droAna3.scaffold_13266 18296409 96 - 19884421 UAAAAACUUUCUCCUUUCCGGAGAC-----ACUUUGUGUACCU------------CAGCCCGUUCAGACUGUUUACCAAAUCACGGCCAAUUAGAACACA---UGAGGUGCGCUAC .........(((((.....))))).-----.....((((((((------------((....((((.((.((....((.......)).)).)).))))...---))))))))))... ( -26.30, z-score = -2.97, R) >droVir3.scaffold_12875 9551520 115 + 20611582 UCUAUACGUUU-CCUGUGUAGAUGUGUGUAUGUGAAUGCUGGGGUAGGUUUCGGGGCGUUGGUGCAUUUAGUAUGCGGCAGCAAGGUUAACUAGUUUCCAGUCUCUCAAAAGCAAU ((((((((...-..))))))))..............((((.(((.((..(..(..((((((.(((((.....))))).)))).((.....)).))..).)..)))))...)))).. ( -31.40, z-score = -0.71, R) >consensus CACAAACUUUCGCCUUCAUUGACGCCUGCAAACCGGUGAGCCCUUCGGAAGCGC_CCACUCAAACAGCCUGGUUAUAAAAUCA_GGCCAAUUAGUGCACG___UGAGAAAGGCGAC ......................(((((........(((..................))).......((((((((....))))).)))......................))))).. ( -6.28 = -6.61 + 0.33)

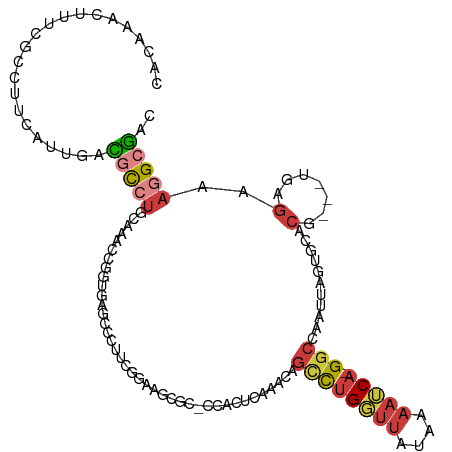

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:08:15 2011