
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 2,333,321 – 2,333,425 |
| Length | 104 |
| Max. P | 0.743522 |

| Location | 2,333,321 – 2,333,425 |
|---|---|
| Length | 104 |
| Sequences | 9 |
| Columns | 109 |
| Reading direction | forward |
| Mean pairwise identity | 72.35 |
| Shannon entropy | 0.59662 |
| G+C content | 0.53779 |
| Mean single sequence MFE | -35.65 |
| Consensus MFE | -11.72 |
| Energy contribution | -12.38 |
| Covariance contribution | 0.66 |
| Combinations/Pair | 1.42 |
| Mean z-score | -1.96 |
| Structure conservation index | 0.33 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.56 |
| SVM RNA-class probability | 0.743522 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 2333321 104 + 23011544 GAAUCGGUGGCAGGAGAUGAACCCAAAGUGCAUUUCAUUGCUCACUGUCGCAGGUCGCAGGCAGCGAUUGCGAUGGCGAUGGAUUCGAAGGAAGCUGGCCAACC----- ....(((((((((((((((.((.....)).)))))).)))).)))))((((((.((((.....))))))))))((((.(.(..(((....))).)).))))...----- ( -38.20, z-score = -1.49, R) >droWil1.scaffold_180708 10254856 79 - 12563649 -------CAGCAUUCGAUGAG-CCAAAGUGCAUUUCAUUGCUCACUGUCACCAAACGGGGAGGGUGGGCGAGUGGGUGAUUGAAGAA---------------------- -------.....((((((.(.-(((............(((((((((.((.((....)).)).))))))))).))).).))))))...---------------------- ( -27.22, z-score = -2.03, R) >droPer1.super_1 4734530 107 - 10282868 GCAGCAUCAGCAGUAGAUGAAACCAAAGUGCAUUUCAUUGCUCACUGUCGCAGGUCCCCAGCAGCGAUU--UACGGCAAUAGAUUCCCAAAAGGAAGAGCUGUCCAAUC (((((.(((((((((((((..((....)).))))).))))))...(((((.((((((......).))))--).))))).....((((.....)))))))))))...... ( -24.90, z-score = -0.59, R) >dp4.chr4_group3 7616252 107 - 11692001 GCAGCAUCAGCAGUAGAUGAAACCAAAGUGCAUUUCAUUGCUCACUGUCGCAGGUCCCCAGCAGCGAUU--UACGGCAAUAGAUUCCCAAAAGGAAGAGCUGUCCAAUC (((((.(((((((((((((..((....)).))))).))))))...(((((.((((((......).))))--).))))).....((((.....)))))))))))...... ( -24.90, z-score = -0.59, R) >droAna3.scaffold_12916 10053733 102 - 16180835 UAUCCAGUGGCAGGAGAUGAACCCAAAGUGCAUUUCAUUGCUCACUGUCGCAGGUCGCAGCUAGCGAUU--UACGGCAAUGGAUUCGGAUUUAGCUCGGAACCC----- ..(((((((((((((((((.((.....)).)))))).)))).)))(((((.(((((((.....))))))--).))))).))))(((.((......)).)))...----- ( -34.50, z-score = -2.11, R) >droEre2.scaffold_4929 2372323 102 + 26641161 GAAUCGGUGGCAGGAGAUGGACCCAAAGUGCAUUUCAUUGCUCACUGUCGCAGGUCGCAGGCAGCGAUU--UGCGACGAUGGAUUCGAAGGAAGCCGGCCAACC----- ..(((((((((((((((((.((.....)).)))))).)))).))))((((((((((((.....))))))--)))))))))((.(((....))).))........----- ( -44.80, z-score = -3.78, R) >droYak2.chr2L 2312800 105 + 22324452 GAAUCGGUGGCAGGAGAUGGACCCAAAGUGCAUUUCAUUGCUCACAGUCGCAGGUCGCAGGCAGCGCGAUUUGGGGCGAUGGAUUCGAAGGAAGCUGGCCAAGCC---- (((((.(((((((((((((.((.....)).)))))).)))).))).(((.((((((((.......)))))))).)))....))))).......(((.....))).---- ( -39.90, z-score = -1.85, R) >droSec1.super_5 504862 106 + 5866729 GAAUCGGUGGCAGGAGAUGAACCCAAAGUGCAUUUCAUUGCUCACUGUCGCAGGUCGCAGGCAGCGAUU--UGCGGCGAUGGAUUCGAAGGAAGCUGGCCAGCCAACC- (((((((((((((((((((.((.....)).)))))).)))).))))((((((((((((.....))))))--))))))....))))).........(((....)))...- ( -43.20, z-score = -2.59, R) >droSim1.chr2L 2290714 106 + 22036055 GAAUCGGUGGCAGGAGAUGAACCCAAAGUGCAUUUCAUUGCUCACUGUCGCAGGUCGCAGGCAGCGAUU--UGCGGCGAUGGAUUCGAAGGAAGCUGGCCAGCCAACC- (((((((((((((((((((.((.....)).)))))).)))).))))((((((((((((.....))))))--))))))....))))).........(((....)))...- ( -43.20, z-score = -2.59, R) >consensus GAAUCGGUGGCAGGAGAUGAACCCAAAGUGCAUUUCAUUGCUCACUGUCGCAGGUCGCAGGCAGCGAUU__UACGGCGAUGGAUUCGAAGGAAGCUGGCCAGCC_____ ....(((((((((..((((...........))))...)))))...(((((..((((((.....))))))....)))))...............))))............ (-11.72 = -12.38 + 0.66)


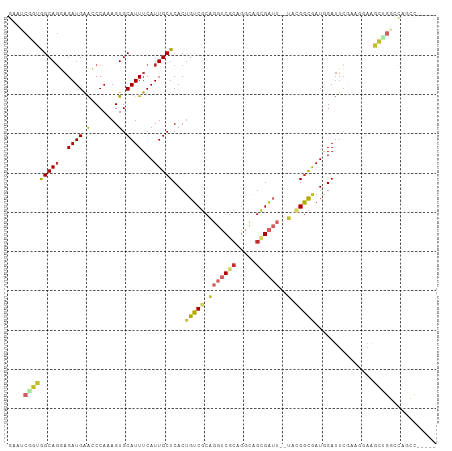
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:10:58 2011