
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 4,246,113 – 4,246,204 |
| Length | 91 |
| Max. P | 0.754893 |

| Location | 4,246,113 – 4,246,204 |
|---|---|
| Length | 91 |
| Sequences | 7 |
| Columns | 115 |
| Reading direction | forward |
| Mean pairwise identity | 61.93 |
| Shannon entropy | 0.67733 |
| G+C content | 0.56826 |
| Mean single sequence MFE | -25.67 |
| Consensus MFE | -10.54 |
| Energy contribution | -10.86 |
| Covariance contribution | 0.32 |
| Combinations/Pair | 1.67 |
| Mean z-score | -1.18 |
| Structure conservation index | 0.41 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.59 |
| SVM RNA-class probability | 0.754893 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
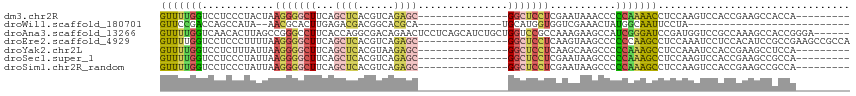
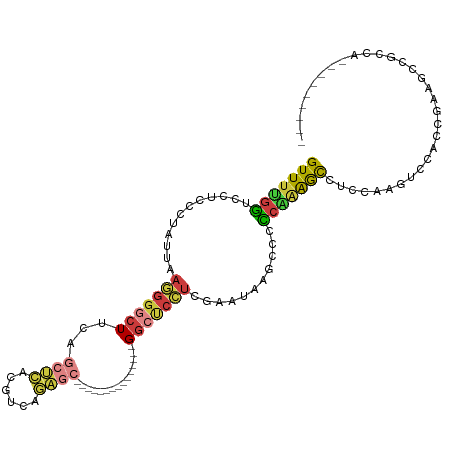
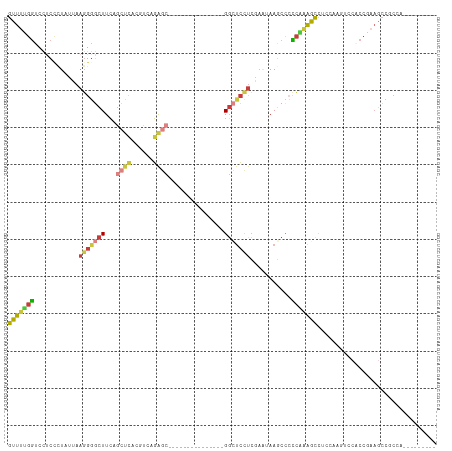
>dm3.chr2R 4246113 91 + 21146708 GUUUUGGUCCUCCCUACUAAGGGGCUUCAGCUCACGUCAGAGC---------------GGCUCCUCGAAUAAACCCCCAAAACCUCCAAGUCCACCGAAGCCACCA--------- ....((((....(((....)))((((((.((((......))))---------------((((..........................))))....))))))))))--------- ( -21.47, z-score = -1.13, R) >droWil1.scaffold_180701 676607 72 + 3904529 GUUCCGACCAGCCAUA--AACGCACUUGAGACGACGGCACGCA--------------UGCAUGGUGGUCGAAACUAUGGCAAUUCCUA--------------------------- (((.((((((.((((.--..(((......).))...(((....--------------))))))))))))).)))..............--------------------------- ( -17.80, z-score = -0.22, R) >droAna3.scaffold_13266 18135252 109 - 19884421 GUUUUGGUCAACACUUAGCCGGGCCUUCACCAGGCGACAGAACUCCUCAGCAUCUGCUGGUCCGCCAAAGAAGCCAUCGGGAUCCGAUGGUCCGCCAAAGCCACCGGGA------ ((((((((.....(((.(.(((((((.....))))...........(((((....))))).))).).)))..((((((((...))))))))..))))))))........------ ( -36.30, z-score = -0.39, R) >droEre2.scaffold_4929 8254090 100 - 26641161 GUUUUGGUCCUCCCUUUUAAGGGGCUUCAGCUCACGUCAGAGC---------------GGCUCCUCAAGUAAGCCCCCCAAGCCUCCAAAUCCUCCACAUCCGCCGAAGCCGCCA ((((((((....(((....)))(((((..((((......))))---------------((((.........))))....)))))..................))))))))..... ( -26.10, z-score = -1.54, R) >droYak2.chr2L 16901808 91 + 22324452 GUUUUGGUCCUCUUUAUUAAGGGGCUUCAGCUCACGUAAGAGC---------------GGCUCCUCAAGCAAGCCCCCAAAGCCUCCAAAUCCACCGAAGCCUCCA--------- ((((((((............(((((((..((((......))))---------------.(((.....))))))))))................)))))))).....--------- ( -25.99, z-score = -2.07, R) >droSec1.super_1 1892151 91 + 14215200 GUUUUGGUCCUCCCUAUUAAGGGGCUUCAGCUCACGUCAGAGC---------------GGCUCCUCGAAUAAGCCCCCAAAGCCUCCAAGUCCACCGAAGCCGCCA--------- ((((((((....(((....)))(((((..((((......))))---------------((((.........))))....))))).........)))))))).....--------- ( -26.00, z-score = -1.46, R) >droSim1.chr2R_random 1382531 91 + 2996586 GUUUUGGUCCUCCCUAUUAAGGGGCUUCAGCUCACGUCAGAGC---------------GGCUCCUCGAAUAAGCCCCCAAAGCCUCCAAGUCCACCGAAGCCGCCA--------- ((((((((....(((....)))(((((..((((......))))---------------((((.........))))....))))).........)))))))).....--------- ( -26.00, z-score = -1.46, R) >consensus GUUUUGGUCCUCCCUAUUAAGGGGCUUCAGCUCACGUCAGAGC_______________GGCUCCUCGAAUAAGCCCCCAAAGCCUCCAAGUCCACCGAAGCCGCCA_________ (((((((............(((((((...((((......))))...............)))))))...........)))))))................................ (-10.54 = -10.86 + 0.32)
| Location | 4,246,113 – 4,246,204 |
|---|---|
| Length | 91 |
| Sequences | 7 |
| Columns | 115 |
| Reading direction | reverse |
| Mean pairwise identity | 61.93 |
| Shannon entropy | 0.67733 |
| G+C content | 0.56826 |
| Mean single sequence MFE | -34.13 |
| Consensus MFE | -12.62 |
| Energy contribution | -13.12 |
| Covariance contribution | 0.50 |
| Combinations/Pair | 1.67 |
| Mean z-score | -1.29 |
| Structure conservation index | 0.37 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.35 |
| SVM RNA-class probability | 0.655744 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
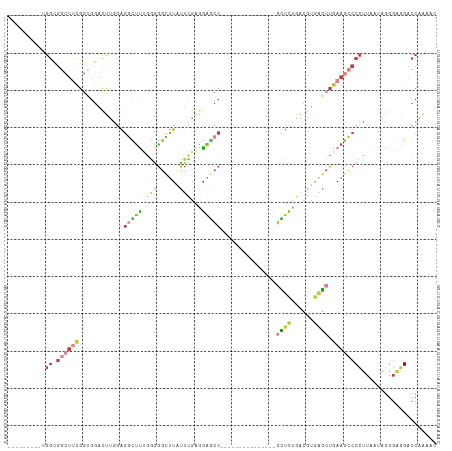
>dm3.chr2R 4246113 91 - 21146708 ---------UGGUGGCUUCGGUGGACUUGGAGGUUUUGGGGGUUUAUUCGAGGAGCC---------------GCUCUGACGUGAGCUGAAGCCCCUUAGUAGGGAGGACCAAAAC ---------(((((((((((((..(((..((((((((..(((....)))..))))))---------------..))..).))..)))))))))((((......)))))))).... ( -32.30, z-score = -1.00, R) >droWil1.scaffold_180701 676607 72 - 3904529 ---------------------------UAGGAAUUGCCAUAGUUUCGACCACCAUGCA--------------UGCGUGCCGUCGUCUCAAGUGCGUU--UAUGGCUGGUCGGAAC ---------------------------..((.....))...(((((((((((((((..--------------.(((..(...........)..))).--))))).)))))))))) ( -26.10, z-score = -2.29, R) >droAna3.scaffold_13266 18135252 109 + 19884421 ------UCCCGGUGGCUUUGGCGGACCAUCGGAUCCCGAUGGCUUCUUUGGCGGACCAGCAGAUGCUGAGGAGUUCUGUCGCCUGGUGAAGGCCCGGCUAAGUGUUGACCAAAAC ------....(((((((((.((((((((((((...))))))).)))...(((((..((((....)))).((....)).)))))..)).))))))((((.....)))))))..... ( -40.40, z-score = -0.35, R) >droEre2.scaffold_4929 8254090 100 + 26641161 UGGCGGCUUCGGCGGAUGUGGAGGAUUUGGAGGCUUGGGGGGCUUACUUGAGGAGCC---------------GCUCUGACGUGAGCUGAAGCCCCUUAAAAGGGAGGACCAAAAC (((.(((((((((..((((((((....(((...((..((.......))..))...))---------------))))).))))..)))))))))((((......)))).))).... ( -37.10, z-score = -1.26, R) >droYak2.chr2L 16901808 91 - 22324452 ---------UGGAGGCUUCGGUGGAUUUGGAGGCUUUGGGGGCUUGCUUGAGGAGCC---------------GCUCUUACGUGAGCUGAAGCCCCUUAAUAAAGAGGACCAAAAC ---------(((.(((((((...........((((((..(((....)))..))))))---------------((((......)))))))))))((((......)))).))).... ( -33.00, z-score = -1.39, R) >droSec1.super_1 1892151 91 - 14215200 ---------UGGCGGCUUCGGUGGACUUGGAGGCUUUGGGGGCUUAUUCGAGGAGCC---------------GCUCUGACGUGAGCUGAAGCCCCUUAAUAGGGAGGACCAAAAC ---------(((.(((((((((...((((((((((.....))))..))))))..)))---------------((((......)))).))))))((((......)))).))).... ( -35.00, z-score = -1.36, R) >droSim1.chr2R_random 1382531 91 - 2996586 ---------UGGCGGCUUCGGUGGACUUGGAGGCUUUGGGGGCUUAUUCGAGGAGCC---------------GCUCUGACGUGAGCUGAAGCCCCUUAAUAGGGAGGACCAAAAC ---------(((.(((((((((...((((((((((.....))))..))))))..)))---------------((((......)))).))))))((((......)))).))).... ( -35.00, z-score = -1.36, R) >consensus _________UGGCGGCUUCGGUGGACUUGGAGGCUUUGGGGGCUUAUUCGAGGAGCC_______________GCUCUGACGUGAGCUGAAGCCCCUUAAUAGGGAGGACCAAAAC ..........((.(((((((((........(((((.....)))))......(((((................))))).......))))))))))).................... (-12.62 = -13.12 + 0.50)


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:07:46 2011