
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 3,790,012 – 3,790,152 |
| Length | 140 |
| Max. P | 0.571485 |

| Location | 3,790,012 – 3,790,152 |
|---|---|
| Length | 140 |
| Sequences | 6 |
| Columns | 162 |
| Reading direction | forward |
| Mean pairwise identity | 78.64 |
| Shannon entropy | 0.37674 |
| G+C content | 0.41335 |
| Mean single sequence MFE | -36.62 |
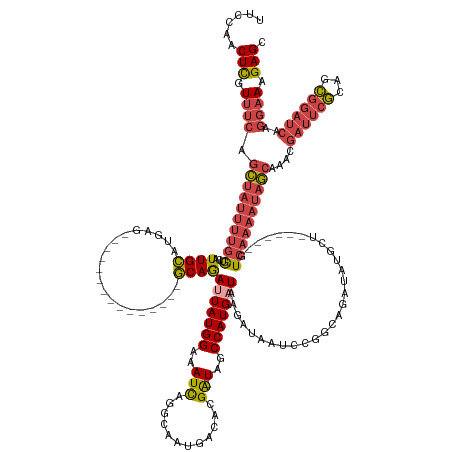
| Consensus MFE | -19.86 |
| Energy contribution | -21.89 |
| Covariance contribution | 2.03 |
| Combinations/Pair | 1.26 |
| Mean z-score | -1.80 |
| Structure conservation index | 0.54 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.16 |
| SVM RNA-class probability | 0.571485 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
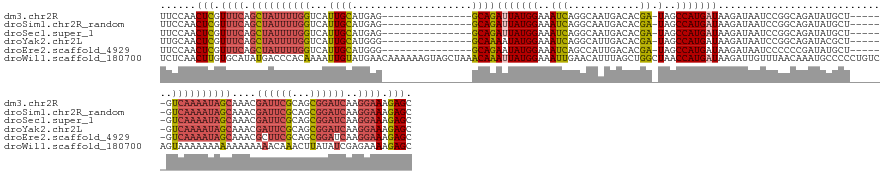
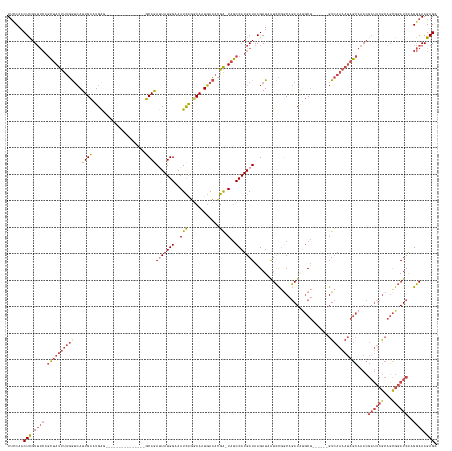
>dm3.chr2R 3790012 140 + 21146708 UUCCAACUCGUUUCAGCUAUUUUGGUCAUUGCAUGAG---------------GCAGAUUAUGGAAAUCAGGCAAUGACACGA-UAGCCAUGAUAAGAUAAUCCGGCAGAUAUGCU------GUCAAAAUAGCAAACGAUUCGCAGCGGAUCAAGGAAAGAGC ......(((.((((.(((((((((((((((((.(((.---------------.(........)...))).))))))))....-...................(((((....))))------).)))))))))....((((((...))))))..)))).))). ( -41.40, z-score = -2.69, R) >droSim1.chr2R_random 1023701 140 + 2996586 UUCCAACUCGUUUCAGCUAUUUUGGUCAUUGCAUGAG---------------GCAGAUUAUGGAAAUCAGGCAAUGACACGA-UAGCCAUGAUAAGAUAAUCCGGCAGAUAUGCU------GUCAAAAUAGCAAACGAUUCGCAGCGGAUCAAGGAAAGAGC ......(((.((((.(((((((((((((((((.(((.---------------.(........)...))).))))))))....-...................(((((....))))------).)))))))))....((((((...))))))..)))).))). ( -41.40, z-score = -2.69, R) >droSec1.super_1 1426522 140 + 14215200 UUCCAACUCGUUUCAGCUAUUUUGGUCAUUGCAUGAG---------------GCAGAUUAUGGAAAUCAGGCAAUGACACGA-UAGCCAUGAUAAGAUAAUCCGGCAGAUAUGCU------GUCAAAAUAGCAAACGAUUCGCAGCGGAUCAAGGAAAGAGC ......(((.((((.(((((((((((((((((.(((.---------------.(........)...))).))))))))....-...................(((((....))))------).)))))))))....((((((...))))))..)))).))). ( -41.40, z-score = -2.69, R) >droYak2.chr2L 16447103 140 + 22324452 UUGCAACUCGUUUCAGCUAUUUUGGUCAUUGCAUGGG---------------GCAAAAUAUGGAAAUCAGGCAUUGACACGA-UAGCCAUGAUAAGAUAAUCCGGCAGAUACGCU------GUCAAAAUAGCAAACGAUUCGCAGCGGAUCAAGGAAAGAGC ......(((.((((.((((((((((((.((((.....---------------)))).........(((((((((((...)))-).))).))))..)))....((((......)))------).)))))))))....((((((...))))))..)))).))). ( -37.60, z-score = -1.64, R) >droEre2.scaffold_4929 7803972 140 - 26641161 UUCCAACUCGUUUCAGCUAUUUUGGUCAUUGCAUGGG---------------GCAGAAUAUGGAAAUCAGCCAUUGACACGA-UAGCCAUGAUAAGAUAAUCCCCCCGAUAUGCU------GUCAAAAUAGCAAACGCUUCGCAGCGGAUCAAGGAAAGAGC ......(((.((((.((((((((((.((..(((((((---------------(......((((..(((............))-)..))))(((......))).))))...)))))------)))))))))))...((((....))))......)))).))). ( -35.90, z-score = -0.66, R) >droWil1.scaffold_180700 5521868 162 + 6630534 UCUCAACUUGUGCAUAUGACCCACAAAAUUGUAUGAACAAAAAAGUAGCUAAACAAAUUAUGGAAAUUGAACAUUUAGCUGGCUAACCAUGAUAAGAUUGUUUAACAAAUGCCCCCUGUCAGUAAAAAAAAAAAAAAAACAAACUUAUAUCGAGAAAAGAGC ((((...((((.((((..(.........)..)))).))))...((((((((((......((....))......))))))).))).......(((((.((((((.................................)))))).)))))...))))....... ( -22.01, z-score = -0.40, R) >consensus UUCCAACUCGUUUCAGCUAUUUUGGUCAUUGCAUGAG_______________GCAGAUUAUGGAAAUCAGGCAAUGACACGA_UAGCCAUGAUAAGAUAAUCCGGCAGAUAUGCU______GUCAAAAUAGCAAACGAUUCGCAGCGGAUCAAGGAAAGAGC ......(((.((((.(((((((((((((.(((.(((..............................))).))).)))).......(((..(((......))).))).................)))))))))....((((((...))))))..)))).))). (-19.86 = -21.89 + 2.03)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:06:47 2011