
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 3,591,089 – 3,591,192 |
| Length | 103 |
| Max. P | 0.997612 |

| Location | 3,591,089 – 3,591,192 |
|---|---|
| Length | 103 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 69.46 |
| Shannon entropy | 0.54397 |
| G+C content | 0.43187 |
| Mean single sequence MFE | -29.12 |
| Consensus MFE | -13.91 |
| Energy contribution | -14.83 |
| Covariance contribution | 0.92 |
| Combinations/Pair | 1.24 |
| Mean z-score | -2.59 |
| Structure conservation index | 0.48 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.76 |
| SVM RNA-class probability | 0.995029 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
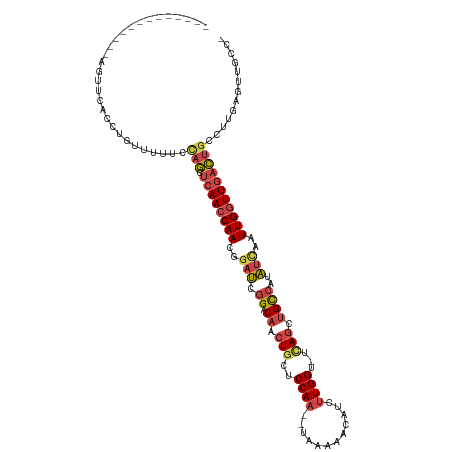
>dm3.chr2R 3591089 103 + 21146708 -------------AGUCCACCCGUUUUUCCAGCUCAACCAACGGAUCGGACAACUGCUCCAA--UAAAAACAUCUUGGU-UAAGCUGCCAUAUUCAUUGCUUGACUGCGUUGAGUUGCC- -------------................((((((((((...(((.(((....))).)))..--............(((-(((((.............))))))))).)))))))))..- ( -26.42, z-score = -2.07, R) >droSim1.chr2R 2314705 103 + 19596830 -------------AGUUCACCUGUAUUUCCAGCUCAACCAACGGAUCGGACAACUGCUCCAA--UAAAAACAUCUUGGU-UCAGCUGCCAUAUCACUUGGUUGACUGCCUUGAGAUGCC- -------------.........((((((((((.((((((((..(((.((.((.(((..((((--..........)))).-.))).))))..)))..)))))))))))....))))))).- ( -29.30, z-score = -2.65, R) >droSec1.super_1 1246507 103 + 14215200 -------------AGUUCACCUGUUUUUCCAGCUCAACCAACGGAUCGGACAACUGCUCCAA--UAAAAACCUCUUGGU-UUAGCUGCCAUAUCAAUUGGUUGACUGCCUUGAAUUGCC- -------------.........((..((((((.((((((((..(((.((.((.(((..((((--..........)))).-.))).))))..)))..)))))))))))....)))..)).- ( -24.70, z-score = -1.73, R) >droYak2.chr2L 16263878 116 + 22324452 UGCCUAUUUGAGAAGUUUAACUGUUUUUCUAGCUCAACCAAUGGAUCGGACAACUGCUCCAA--UGAAAAUAUUUUGGU-UCAUCUGCCCUAUUAAUUGGUUGACUGCCUUAAGUUGCC- .((..(((((((.((.....)).......(((.(((((((((.(((.((.((..((..((((--..........)))).-.))..)).)).))).)))))))))))).))))))).)).- ( -26.70, z-score = -1.68, R) >droEre2.scaffold_4929 7624638 104 - 26641161 ------------UGCUGUGGUCAGUUCAACAGCUCAACCAAUGGAUCGGACAACUGCUCCAU--UGAAAAAAUGUUGGC-UCAGCUGCCUUAUUAAUUGGUUGACUGACUUGAGUUGCA- ------------.(((..(((((((.((((........(((((((.(((....))).)))))--))....((((..(((-......))).)))).....)))))))))))..)))....- ( -32.10, z-score = -1.90, R) >droAna3.scaffold_13266 10096698 102 + 19884421 ------------------CACUAUUUUCAAAAGACAACCAAAGACAAUGACUGCUGUUCCAAGGUUCUAUUAUCUUGGGAUCAGUUGUCUUUACCUUUGGUUAGUCCUCUGAAAUUUCCC ------------------......(((((...(((((((((((.....(((.(((((((((((((......))))))))).)))).))).....)))))))).)))...)))))...... ( -35.50, z-score = -5.52, R) >consensus _____________AGUUCACCUGUUUUUCCAGCUCAACCAACGGAUCGGACAACUGCUCCAA__UAAAAACAUCUUGGU_UCAGCUGCCAUAUCAAUUGGUUGACUGCCUUGAGUUGCC_ .............................(((.((((((((..(((.((.((.(((..((((............))))...))).))))..)))..)))))))))))............. (-13.91 = -14.83 + 0.92)


| Location | 3,591,089 – 3,591,192 |
|---|---|
| Length | 103 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 69.46 |
| Shannon entropy | 0.54397 |
| G+C content | 0.43187 |
| Mean single sequence MFE | -31.38 |
| Consensus MFE | -21.65 |
| Energy contribution | -21.52 |
| Covariance contribution | -0.13 |
| Combinations/Pair | 1.35 |
| Mean z-score | -2.18 |
| Structure conservation index | 0.69 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 3.14 |
| SVM RNA-class probability | 0.997612 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
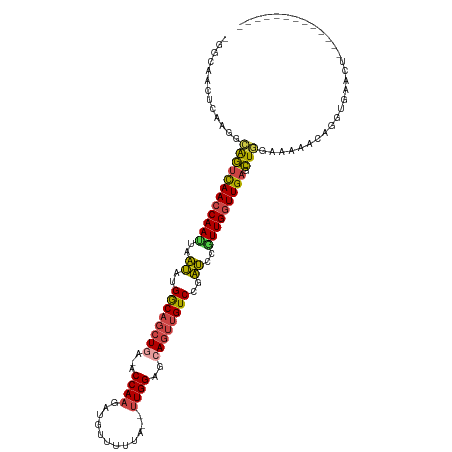
>dm3.chr2R 3591089 103 - 21146708 -GGCAACUCAACGCAGUCAAGCAAUGAAUAUGGCAGCUUA-ACCAAGAUGUUUUUA--UUGGAGCAGUUGUCCGAUCCGUUGGUUGAGCUGGAAAAACGGGUGGACU------------- -(...((((...((......))...........(((((((-(((((((((....))--))(((.(........).))).)))))))))))).......))))...).------------- ( -25.80, z-score = 0.19, R) >droSim1.chr2R 2314705 103 - 19596830 -GGCAUCUCAAGGCAGUCAACCAAGUGAUAUGGCAGCUGA-ACCAAGAUGUUUUUA--UUGGAGCAGUUGUCCGAUCCGUUGGUUGAGCUGGAAAUACAGGUGAACU------------- -..(((((.....(((((((((((..(((..((((((((.-.((((..........--))))..))))))))..)))..)))))))).))).......)))))....------------- ( -34.20, z-score = -2.19, R) >droSec1.super_1 1246507 103 - 14215200 -GGCAAUUCAAGGCAGUCAACCAAUUGAUAUGGCAGCUAA-ACCAAGAGGUUUUUA--UUGGAGCAGUUGUCCGAUCCGUUGGUUGAGCUGGAAAAACAGGUGAACU------------- -.....((((......(((((((((.(((..(((((((..-.((((..........--))))...)))))))..))).))))))))).(((......))).))))..------------- ( -30.00, z-score = -1.54, R) >droYak2.chr2L 16263878 116 - 22324452 -GGCAACUUAAGGCAGUCAACCAAUUAAUAGGGCAGAUGA-ACCAAAAUAUUUUCA--UUGGAGCAGUUGUCCGAUCCAUUGGUUGAGCUAGAAAAACAGUUAAACUUCUCAAAUAGGCA -.((.......(((..(((((((((.....((((((.((.-.((((..........--))))..)).)))))).....))))))))))))...............((........)))). ( -27.30, z-score = -1.45, R) >droEre2.scaffold_4929 7624638 104 + 26641161 -UGCAACUCAAGUCAGUCAACCAAUUAAUAAGGCAGCUGA-GCCAACAUUUUUUCA--AUGGAGCAGUUGUCCGAUCCAUUGGUUGAGCUGUUGAACUGACCACAGCA------------ -...............(((((((((..((..((((((((.-.(((...........--.)))..))))))))..))..)))))))))(((((.(......).))))).------------ ( -33.40, z-score = -2.81, R) >droAna3.scaffold_13266 10096698 102 - 19884421 GGGAAAUUUCAGAGGACUAACCAAAGGUAAAGACAACUGAUCCCAAGAUAAUAGAACCUUGGAACAGCAGUCAUUGUCUUUGGUUGUCUUUUGAAAAUAGUG------------------ ......(((((((((((.((((((((..((.(((..(((...(((((..........)))))..)))..))).))..)))))))))))))))))))......------------------ ( -37.60, z-score = -5.26, R) >consensus _GGCAACUCAAGGCAGUCAACCAAUUAAUAUGGCAGCUGA_ACCAAGAUGUUUUUA__UUGGAGCAGUUGUCCGAUCCGUUGGUUGAGCUGGAAAAACAGGUGAACU_____________ .............((((((((((((..((..((((((((...((((............))))..))))))))..))..))))))))).)))............................. (-21.65 = -21.52 + -0.13)


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:06:13 2011