
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 3,512,422 – 3,512,478 |
| Length | 56 |
| Max. P | 0.897472 |

| Location | 3,512,422 – 3,512,478 |
|---|---|
| Length | 56 |
| Sequences | 7 |
| Columns | 61 |
| Reading direction | forward |
| Mean pairwise identity | 68.49 |
| Shannon entropy | 0.61479 |
| G+C content | 0.52246 |
| Mean single sequence MFE | -12.83 |
| Consensus MFE | -5.85 |
| Energy contribution | -7.10 |
| Covariance contribution | 1.25 |
| Combinations/Pair | 1.38 |
| Mean z-score | -1.45 |
| Structure conservation index | 0.46 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.13 |
| SVM RNA-class probability | 0.897472 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
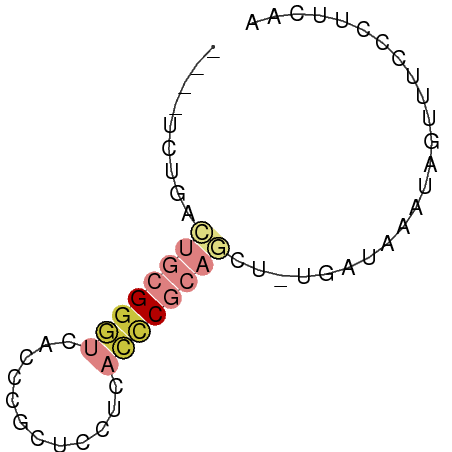
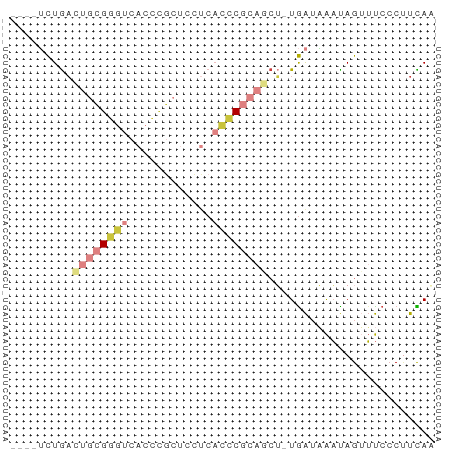
>dm3.chr2R 3512422 56 + 21146708 ----UCUGACUGCGGGUCACCCGCUCCUCACCCGCAGCU-UGAUAAAUAGUUUCCCUUCAA ----((...((((((((............))))))))..-.)).................. ( -13.40, z-score = -2.11, R) >droSim1.chr2R 2237488 56 + 19596830 ----UCUGACUGCGGGUCACCCGCUCCUCACCCGCAACU-UGAUAAAUAGUUUCCCUUCAA ----......(((((((............)))))))...-..................... ( -10.50, z-score = -1.51, R) >droSec1.super_1 1169666 56 + 14215200 ----UCUGACUGCGGGUCACCCGCUCCUCACCCGCAGCU-UGAUAAAUAGAUUCCCUUCAA ----((...((((((((............))))))))..-.)).................. ( -13.40, z-score = -2.06, R) >droYak2.chr2L 16184684 56 + 22324452 ----UCUGACUGCGGGUCACCCGCUUCUCACCCGCAGCU-UGAUAAAUAGUUUCCCUUUAG ----((...((((((((............))))))))..-.)).................. ( -13.40, z-score = -2.05, R) >droEre2.scaffold_4929 7549514 56 - 26641161 ----UCUGACUGCGGGUCACCCGCUCCUCACCCGCAGCU-UGAUAAAUAGUUUCCCUUCAA ----((...((((((((............))))))))..-.)).................. ( -13.40, z-score = -2.11, R) >dp4.chr3 7540167 61 - 19779522 ACUGACUGCUGUGGGCCUACUUCCAGCCCCCUCGCGACGGUGGGUGCCAGAGCCACAUCAA ..(((..(((.((((((((((....((......))...))))))).))).)))....))). ( -20.00, z-score = -0.06, R) >droWil1.scaffold_180569 1211973 56 - 1405142 ----GUUAAAAUUGAUACAUCACCGAGGCAGCCUCCGAG-UAAUUAAUAGUUUACCCUUAA ----......((((((((.((...((((...)))).)))-).))))))............. ( -5.70, z-score = -0.23, R) >consensus ____UCUGACUGCGGGUCACCCGCUCCUCACCCGCAGCU_UGAUAAAUAGUUUCCCUUCAA .........((((((((............))))))))........................ ( -5.85 = -7.10 + 1.25)

| Location | 3,512,422 – 3,512,478 |
|---|---|
| Length | 56 |
| Sequences | 7 |
| Columns | 61 |
| Reading direction | reverse |
| Mean pairwise identity | 68.49 |
| Shannon entropy | 0.61479 |
| G+C content | 0.52246 |
| Mean single sequence MFE | -17.26 |
| Consensus MFE | -7.66 |
| Energy contribution | -8.69 |
| Covariance contribution | 1.03 |
| Combinations/Pair | 1.53 |
| Mean z-score | -1.44 |
| Structure conservation index | 0.44 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.91 |
| SVM RNA-class probability | 0.851222 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
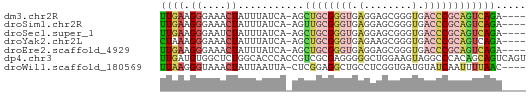
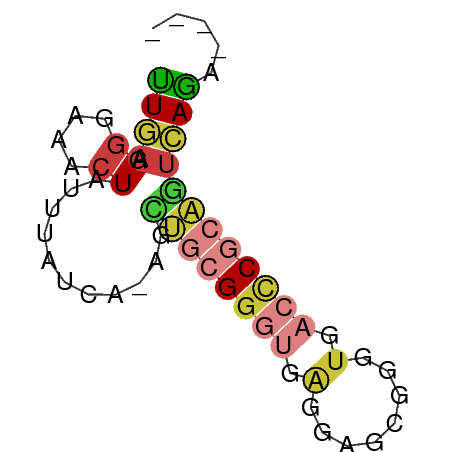
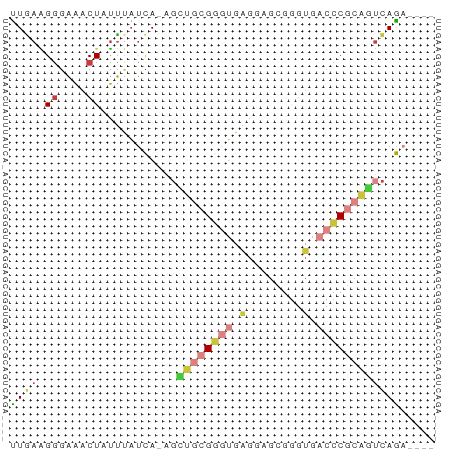
>dm3.chr2R 3512422 56 - 21146708 UUGAAGGGAAACUAUUUAUCA-AGCUGCGGGUGAGGAGCGGGUGACCCGCAGUCAGA---- .....((....)).....((.-.(((((((((.(........).)))))))))..))---- ( -19.30, z-score = -2.39, R) >droSim1.chr2R 2237488 56 - 19596830 UUGAAGGGAAACUAUUUAUCA-AGUUGCGGGUGAGGAGCGGGUGACCCGCAGUCAGA---- ((((.((....))........-..((((((((.(........).)))))))))))).---- ( -16.30, z-score = -1.67, R) >droSec1.super_1 1169666 56 - 14215200 UUGAAGGGAAUCUAUUUAUCA-AGCUGCGGGUGAGGAGCGGGUGACCCGCAGUCAGA---- ..................((.-.(((((((((.(........).)))))))))..))---- ( -16.90, z-score = -1.22, R) >droYak2.chr2L 16184684 56 - 22324452 CUAAAGGGAAACUAUUUAUCA-AGCUGCGGGUGAGAAGCGGGUGACCCGCAGUCAGA---- .....((....)).....((.-.(((((((((.(........).)))))))))..))---- ( -19.30, z-score = -2.60, R) >droEre2.scaffold_4929 7549514 56 + 26641161 UUGAAGGGAAACUAUUUAUCA-AGCUGCGGGUGAGGAGCGGGUGACCCGCAGUCAGA---- .....((....)).....((.-.(((((((((.(........).)))))))))..))---- ( -19.30, z-score = -2.39, R) >dp4.chr3 7540167 61 + 19779522 UUGAUGUGGCUCUGGCACCCACCGUCGCGAGGGGGCUGGAAGUAGGCCCACAGCAGUCAGU (((.(((((..(((.(..(((((.((....)).)).)))..))))..))))).)))..... ( -21.30, z-score = 0.25, R) >droWil1.scaffold_180569 1211973 56 + 1405142 UUAAGGGUAAACUAUUAAUUA-CUCGGAGGCUGCCUCGGUGAUGUAUCAAUUUUAAC---- ....((((((........)))-))).((((...))))....................---- ( -8.40, z-score = -0.04, R) >consensus UUGAAGGGAAACUAUUUAUCA_AGCUGCGGGUGAGGAGCGGGUGACCCGCAGUCAGA____ ((((.((....))...........((((((((.(........).))))))))))))..... ( -7.66 = -8.69 + 1.03)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:06:01 2011