
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 2,922,528 – 2,922,578 |
| Length | 50 |
| Max. P | 0.510134 |

| Location | 2,922,528 – 2,922,578 |
|---|---|
| Length | 50 |
| Sequences | 5 |
| Columns | 64 |
| Reading direction | forward |
| Mean pairwise identity | 59.87 |
| Shannon entropy | 0.61628 |
| G+C content | 0.37700 |
| Mean single sequence MFE | -11.55 |
| Consensus MFE | -3.92 |
| Energy contribution | -5.36 |
| Covariance contribution | 1.44 |
| Combinations/Pair | 1.43 |
| Mean z-score | -1.56 |
| Structure conservation index | 0.34 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.03 |
| SVM RNA-class probability | 0.510134 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 2922528 50 + 21146708 UUUCAUUAACCGAUUUAUAGAUAGAAU-GAAUGAGCGAUCGGUGAGUCGGU------------- ........((((((((((.(((.....-.........))).))))))))))------------- ( -11.94, z-score = -1.81, R) >droSim1.chr2R 1722915 50 + 19596830 UUUCACUCACCGAUUUAUAGAUAGAAU-GAAUGUGCGAUCGGUGAGUCGGC------------- ..(((((((((((((((((........-...)))).))))))))))).)).------------- ( -16.10, z-score = -2.77, R) >droSec1.super_1 590424 50 + 14215200 UUUCACUCACCGAUUUAUAGAUAGAAU-GAAUGUGCGAUCGGUGAGUCGGC------------- ..(((((((((((((((((........-...)))).))))))))))).)).------------- ( -16.10, z-score = -2.77, R) >dp4.chr3 1201802 64 + 19779522 UUUCACUCACAGAUUUACGAGUAUAUUCGUAUACAGAGAUACAGAUACAUAUAGAUAGGUAGAA .....((.((..(((((...((((....((((......))))..))))...)))))..)))).. ( -6.80, z-score = -0.24, R) >droPer1.super_2 1370004 64 + 9036312 UUUCACUCACAGAUUUACGAGUAUAUUCGUAUACAGAGAUACAGAUACAUAUAGAUAGGUAGAA .....((.((..(((((...((((....((((......))))..))))...)))))..)))).. ( -6.80, z-score = -0.24, R) >consensus UUUCACUCACCGAUUUAUAGAUAGAAU_GAAUGCGCGAUCGGUGAGUCGGA_____________ ....(((((((((((.....(((........)))..)))))))))))................. ( -3.92 = -5.36 + 1.44)

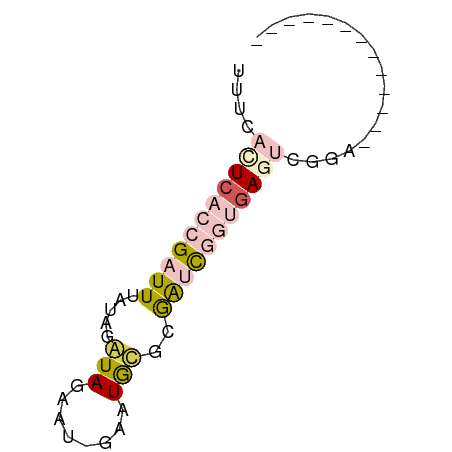

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:04:51 2011