
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 2,913,777 – 2,913,842 |
| Length | 65 |
| Max. P | 0.971355 |

| Location | 2,913,777 – 2,913,842 |
|---|---|
| Length | 65 |
| Sequences | 8 |
| Columns | 65 |
| Reading direction | forward |
| Mean pairwise identity | 67.88 |
| Shannon entropy | 0.69039 |
| G+C content | 0.61296 |
| Mean single sequence MFE | -19.86 |
| Consensus MFE | -10.40 |
| Energy contribution | -10.76 |
| Covariance contribution | 0.36 |
| Combinations/Pair | 1.57 |
| Mean z-score | -1.22 |
| Structure conservation index | 0.52 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.85 |
| SVM RNA-class probability | 0.971355 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 2913777 65 + 21146708 CCACGUUCCCGCCAACUUCCUUGGCGUCUAUCCCUGUCGCCAGAUGGGCUAGCCUGGAGGACGCC ...((((((((((((.....)))))(.(((.(((.(((....)))))).))).).)).))))).. ( -22.50, z-score = -1.21, R) >droSim1.chr2R 1713394 65 + 19596830 CCCCUUUCCCGCCAACUUCCUUGGCGUCUAUUCCUGUCGCCAGAUGGGCUAGCCUGGAGGACGCC ..........((...(((((((((((.(.......).))))))..(((....))))))))..)). ( -18.80, z-score = -0.20, R) >droSec1.super_1 579103 65 + 14215200 CCCCAUUCCCGCCAACUUCAUUGGCGUCUAUUCCUGUCGCCAGAUGGGCUAGCCUGGAGGACGCC (((((....((((((.....)))))).(((..((((((....)))))).)))..))).))..... ( -18.80, z-score = -0.28, R) >droYak2.chr2L 15626982 65 + 22324452 CCACGCUGCCGCCAAUUUCCUUGGCGUCUAUCCCUGUCGCCAGACGGGCUAGCCCGGAGGACGCC ....((....(((((.....)))))((((.(((..(((....)))(((....)))))))))))). ( -23.80, z-score = -1.09, R) >droEre2.scaffold_4929 6989549 65 - 26641161 CCACGCUGCCGACAAUUUCCUUGGCGUCUAUCCCUGUCGCCAGAUGGGCUAGCCUGGAGGACGCC (((.((((((((.....)).((((((.(.......).))))))...))).))).)))........ ( -20.10, z-score = -0.24, R) >droAna3.scaffold_13266 3155885 64 + 19884421 -CCCAAUCCUAGUAAUUUUCAUGGCGUCUAUCUGCGACGCCAGACAGGCUAGUCGUAGGGACGCC -.....................(((((((..((((((((((.....)))..)))))))))))))) ( -24.90, z-score = -3.58, R) >droVir3.scaffold_10324 1054848 62 + 1288806 ---CCCCUUUUGCUCCUACUGCUGUCGUAUUUCCUGACGCCAGCCAGACUAGUCUGGCGGACGCC ---........(((((....((.((((.......))))))..((((((....))))))))).)). ( -19.40, z-score = -2.47, R) >droMoj3.scaffold_6496 21882359 50 - 26866924 ---------------CUCCCUAUGCCUCGGCUACUGUCGCCAACUAGACUAGUCUAUUGGACGCC ---------------........((..((((....))))((((.((((....))))))))..)). ( -10.60, z-score = -0.73, R) >consensus CCACGCUCCCGCCAACUUCCUUGGCGUCUAUCCCUGUCGCCAGAUGGGCUAGCCUGGAGGACGCC ......................(((((((....(((....))).((((....))))..))))))) (-10.40 = -10.76 + 0.36)


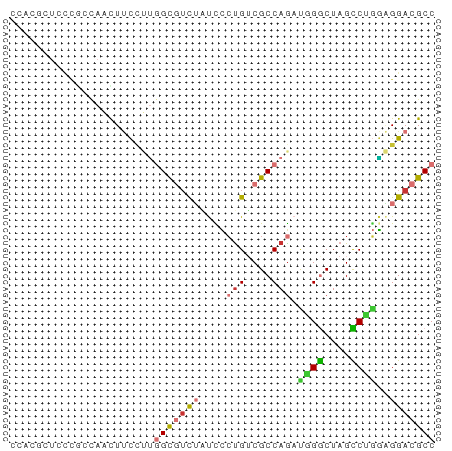
| Location | 2,913,777 – 2,913,842 |
|---|---|
| Length | 65 |
| Sequences | 8 |
| Columns | 65 |
| Reading direction | reverse |
| Mean pairwise identity | 67.88 |
| Shannon entropy | 0.69039 |
| G+C content | 0.61296 |
| Mean single sequence MFE | -20.96 |
| Consensus MFE | -11.84 |
| Energy contribution | -12.20 |
| Covariance contribution | 0.36 |
| Combinations/Pair | 1.46 |
| Mean z-score | -0.59 |
| Structure conservation index | 0.56 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.93 |
| SVM RNA-class probability | 0.856267 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 2913777 65 - 21146708 GGCGUCCUCCAGGCUAGCCCAUCUGGCGACAGGGAUAGACGCCAAGGAAGUUGGCGGGAACGUGG ((((((.(((.((....))...(((....))))))..))))))..........(((....))).. ( -23.70, z-score = -0.53, R) >droSim1.chr2R 1713394 65 - 19596830 GGCGUCCUCCAGGCUAGCCCAUCUGGCGACAGGAAUAGACGCCAAGGAAGUUGGCGGGAAAGGGG (((.(.....).)))..(((.((((....))))......((((((.....)))))).....))). ( -19.70, z-score = 0.59, R) >droSec1.super_1 579103 65 - 14215200 GGCGUCCUCCAGGCUAGCCCAUCUGGCGACAGGAAUAGACGCCAAUGAAGUUGGCGGGAAUGGGG ......(((((......(((.((((....)))).......((((((...)))))))))..))))) ( -22.00, z-score = -0.31, R) >droYak2.chr2L 15626982 65 - 22324452 GGCGUCCUCCGGGCUAGCCCGUCUGGCGACAGGGAUAGACGCCAAGGAAAUUGGCGGCAGCGUGG .((((((..((((....)))).(((....)))))))...((((((.....))))))...)).... ( -26.60, z-score = -0.65, R) >droEre2.scaffold_4929 6989549 65 + 26641161 GGCGUCCUCCAGGCUAGCCCAUCUGGCGACAGGGAUAGACGCCAAGGAAAUUGUCGGCAGCGUGG ........(((.(((.(((..((((....))))(((((...(....)...))))))))))).))) ( -20.90, z-score = 0.16, R) >droAna3.scaffold_13266 3155885 64 - 19884421 GGCGUCCCUACGACUAGCCUGUCUGGCGUCGCAGAUAGACGCCAUGAAAAUUACUAGGAUUGGG- ((((((.((.((((..(((.....))))))).))...)))))).....................- ( -21.00, z-score = -1.72, R) >droVir3.scaffold_10324 1054848 62 - 1288806 GGCGUCCGCCAGACUAGUCUGGCUGGCGUCAGGAAAUACGACAGCAGUAGGAGCAAAAGGGG--- ((((((.((((((....)))))).)))))).............((.......))........--- ( -21.80, z-score = -2.00, R) >droMoj3.scaffold_6496 21882359 50 + 26866924 GGCGUCCAAUAGACUAGUCUAGUUGGCGACAGUAGCCGAGGCAUAGGGAG--------------- .((.((...((((....))))...(((.......))))).))........--------------- ( -12.00, z-score = -0.25, R) >consensus GGCGUCCUCCAGGCUAGCCCAUCUGGCGACAGGAAUAGACGCCAAGGAAGUUGGCGGGAACGGGG ((((((.(((.(....(((.....)))..).)))...))))))...................... (-11.84 = -12.20 + 0.36)


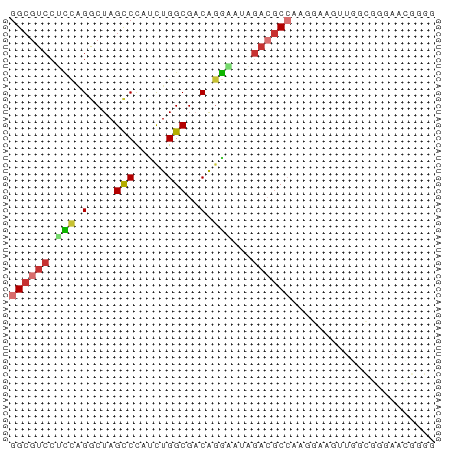
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:04:50 2011