| Sequence ID | dm3.chr2R |
|---|---|
| Location | 2,664,924 – 2,665,009 |
| Length | 85 |
| Max. P | 0.961466 |

| Location | 2,664,924 – 2,665,009 |
|---|---|
| Length | 85 |
| Sequences | 7 |
| Columns | 91 |
| Reading direction | reverse |
| Mean pairwise identity | 68.77 |
| Shannon entropy | 0.63249 |
| G+C content | 0.62284 |
| Mean single sequence MFE | -36.03 |
| Consensus MFE | -15.40 |
| Energy contribution | -18.30 |
| Covariance contribution | 2.90 |
| Combinations/Pair | 1.36 |
| Mean z-score | -1.81 |
| Structure conservation index | 0.43 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.69 |
| SVM RNA-class probability | 0.961466 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
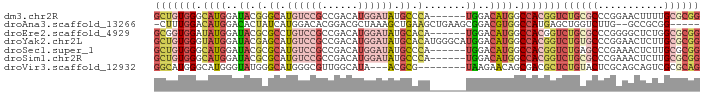
Download alignment: ClustalW | MAF
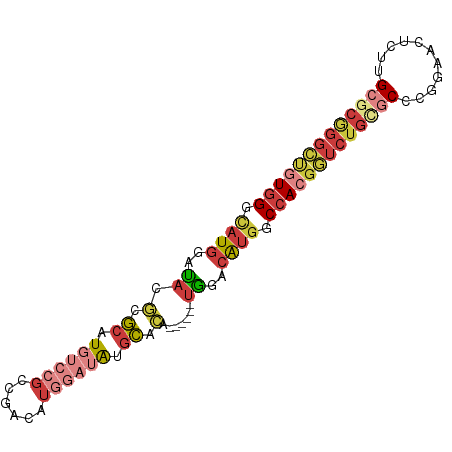
>dm3.chr2R 2664924 85 - 21146708 GCUGUGGGCAUGGAUACGGGCAUGUCCGCCGACAUGGAUAUGCCCA------UGGACAUGGCCACGGUCUGCGCCCGGAACUUUUGCGCGG (((((((.((((..((.(((((((((((......))))))))))).------))..)))).)))))))((((((...........)))))) ( -46.00, z-score = -4.12, R) >droAna3.scaffold_13266 1554628 83 + 19884421 -CUUUGGACAUGGACACUAUCAUGGACACGGACGCUAAAGCUGAAGCUGAAGCGGACGUGGCCAUGAGCUGGUCUUG--GCCGCGG----- -((.(((.((.((((....((((((.((((..((((..(((....)))..))))..)))).))))))....))))))--.))).))----- ( -37.20, z-score = -5.01, R) >droEre2.scaffold_4929 6759862 85 + 26641161 GCGGUGGAUAUGGAUACGCGCCUGUCCGCCGACAUGGAUAUGCACA------UGGACAUGGCCACGGUCUGCGCCCGGGGCUCUGGCGCGG ((((((((((.((.......))))))))))..((((..(((....)------))..))))((((.(((((.......))))).)))))).. ( -33.70, z-score = 0.47, R) >droYak2.chr2L 15400180 91 - 22324452 GCUGUGGGUAUGGAUACGAGCAUGUCCGCCGACAUGGAUAUGCACAUGGGCAUGGACAUGGCCACGGUCUGUGCCCGGAACUCUUGCGCGG ((...((((..........(((((((((......)))))))))...(((((((((((.((....)))))))))))))..))))..)).... ( -37.00, z-score = -1.13, R) >droSec1.super_1 337025 85 - 14215200 GCUGUGGGCAUGGAUACGCGCAUGUCCGCCGACAUGGAUAUGCCCA------UGGACAUGGCCACGGUCUGAGCCCGAAACUCUUGCGCGG (((((((.((((..((.(.(((((((((......))))))))).).------))..)))).)))))))....((.(((.....))).)).. ( -34.30, z-score = -1.28, R) >droSim1.chr2R 1478891 85 - 19596830 GCUGUGGGCAUGGAUACGCGCAUGUCCGCCGACAUGGAUAUGCCCA------UGGACAUGGCCACGGUCUGCGCCCGAAACUCUUGCGCGG (((((((.((((..((.(.(((((((((......))))))))).).------))..)))).)))))))((((((..(.....)..)))))) ( -39.80, z-score = -2.59, R) >droVir3.scaffold_12932 17984 80 + 2102469 GGCAUGGGCAUGGGUAUGGGCAUGGGCGUUGGCAUA---ACGCG--------UAAGAACAGCGACGCUCUGUACUCGCAGCAGUCGCGCAG .......((.......((..(.((.((((((...))---)))).--------)).)..))(((((...((((....))))..))))))).. ( -24.20, z-score = 0.97, R) >consensus GCUGUGGGCAUGGAUACGCGCAUGUCCGCCGACAUGGAUAUGCACA______UGGACAUGGCCACGGUCUGCGCCCGGAACUCUUGCGCGG (((((((.((((.......((.((((((......)))))).)).(((....)))..)))).)))))))((((((...........)))))) (-15.40 = -18.30 + 2.90)

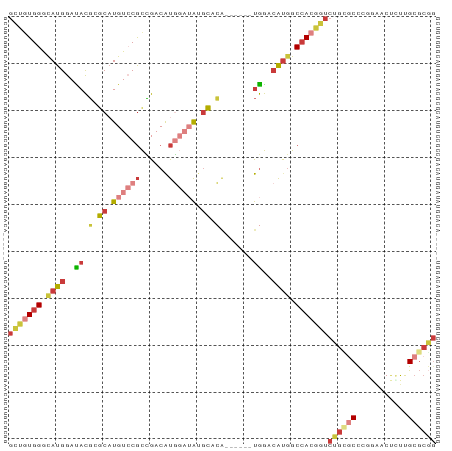
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:04:01 2011