
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 2,449,352 – 2,449,456 |
| Length | 104 |
| Max. P | 0.587533 |

| Location | 2,449,352 – 2,449,456 |
|---|---|
| Length | 104 |
| Sequences | 8 |
| Columns | 105 |
| Reading direction | forward |
| Mean pairwise identity | 64.95 |
| Shannon entropy | 0.69414 |
| G+C content | 0.44311 |
| Mean single sequence MFE | -25.10 |
| Consensus MFE | -10.52 |
| Energy contribution | -9.69 |
| Covariance contribution | -0.83 |
| Combinations/Pair | 1.69 |
| Mean z-score | -0.87 |
| Structure conservation index | 0.42 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.19 |
| SVM RNA-class probability | 0.587533 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 2449352 104 + 21146708 UUUCUCCAUUCCCAAAAAAAA-GUUUGAGAAGAAAAAAGCUAAAAACCGCAGACAGAAUCCUCUGGACUCGAGGACUGGGAACACAAAAAUUCGGCGUCUGCUGC (((((....((.((((.....-.)))).))))))).............((((((.((((((((.......))))).((....))......)))...))))))... ( -20.60, z-score = 0.10, R) >droSim1.chr2R 1276912 104 + 19596830 UUUCUCCAGUCCCAAAAAAAA-GUUUGAGAAGAAAAAAGCUAAAAACCGCAGACAGAAUCCUCUGGACUCGAGGACUGGGAACACAAAAAUUCGGCGUCUGCUGC .(((.(((((((.........-..(((((.........((........))...((((....))))..)))))))))))))))..........((((....)))). ( -24.40, z-score = -0.82, R) >droSec1.super_1 118985 103 + 14215200 UUUCUCCAGUCCCAAAAAAA--GUUUGAGAAGAAAAAAGCUAAAAACCGCAGACAGAAUCCUCUGGACUCGAGGACUGGGAACACAAAAAUUCGGCGUCUGCUGC .(((.(((((((........--..(((((.........((........))...((((....))))..)))))))))))))))..........((((....)))). ( -24.50, z-score = -0.89, R) >droYak2.chr2L 15203687 105 + 22324452 UUACUCCAGUCCCAAAAAAAAUGUUUGAGAAGAAAAAAGCUGAAUACCGCAGACAGAAUCCUCUAAACUGGGGGACUGGGAACACAAAAAUUUAGAGUCUGCUGC ....(((((((((........((((((.(..................).))))))....((........)))))))))))......................... ( -22.17, z-score = -0.66, R) >droEre2.scaffold_4929 6551118 102 - 26641161 UUACUCCAGUGCCAAAAAAA--GUUGGCGAAGAAAAAAGCUGAAAACCGCAGACAGAAUCCUCCGGCCUGCGG-ACUGGGAACACAAAAAUUCGGAGUCUGCUGC ......(((((((((.....--.)))))..........))))......((((.((((.(((.((((((...))-.))))(((........)))))).)))))))) ( -27.20, z-score = -1.04, R) >droAna3.scaffold_13266 1296385 102 - 19884421 -UUUUUUGUUCACUCAAAAAU-GCUAAAGAAGAAAAA-GCUUAAAACCGCAGACAGAAUCCUGAAGCUCGAAGGGCUGGGAACGUAAAAAUUCGCCGUCUGCUGC -.((((((......)))))).-(((...........)-))........((((((....((((..((((.....)))))))).((........))..))))))... ( -21.30, z-score = 0.12, R) >dp4.chr3 18156336 102 + 19779522 ---UUUUUUUCCUGCUGGAAUACCUCGUAAUCGAGGAGAGUCCAAGGGGUAUGCUGUGUCCAAGGAAGUCUUGCGUUAUACCCGCAAAUCUUAGGCAUAUUUUAC ---.......(((..((((...(((((....)))))....)))).)))(((((((..........(((..(((((.......)))))..))).)))))))..... ( -28.80, z-score = -1.47, R) >droPer1.super_34 98863 102 + 916997 ---UUUUUUUCCUGCUGGAAUACCUCGUAAUCGAGGAGAGUCCACGGCGUAUGCGGUGCCCAAGGAAGUCUUGGGUUAUACCCGCAAAUCUUAGACAUAUUUUAC ---........(((.((((...(((((....)))))....)))))))....(((((.(((((((.....))))))).....)))))................... ( -31.80, z-score = -2.29, R) >consensus UUUCUCCAGUCCCAAAAAAAA_GUUUGAGAAGAAAAAAGCUAAAAACCGCAGACAGAAUCCUCUGGACUCGAGGACUGGGAACACAAAAAUUCGGCGUCUGCUGC ................................................((((((....(((((.......))))).((....))............))))))... (-10.52 = -9.69 + -0.83)

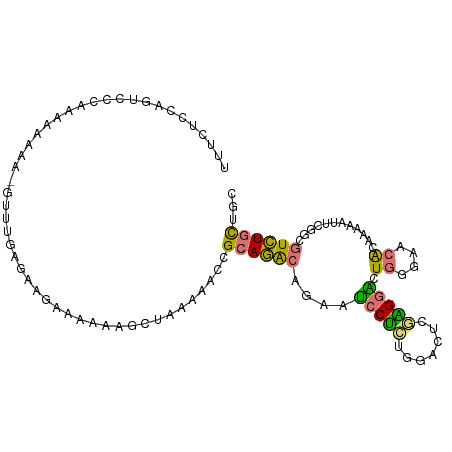

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:03:41 2011