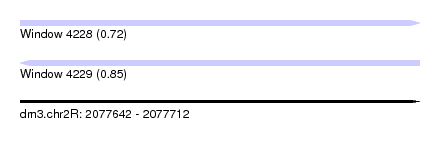

| Sequence ID | dm3.chr2R |
|---|---|
| Location | 2,077,642 – 2,077,712 |
| Length | 70 |
| Max. P | 0.854773 |

| Location | 2,077,642 – 2,077,712 |
|---|---|
| Length | 70 |
| Sequences | 13 |
| Columns | 73 |
| Reading direction | forward |
| Mean pairwise identity | 94.22 |
| Shannon entropy | 0.12272 |
| G+C content | 0.57979 |
| Mean single sequence MFE | -20.12 |
| Consensus MFE | -18.64 |
| Energy contribution | -18.57 |
| Covariance contribution | -0.07 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.32 |
| Structure conservation index | 0.93 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.49 |
| SVM RNA-class probability | 0.717350 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 2077642 70 + 21146708 GUGGACUCGAACCACCAACCUUUCGGUUAACAGCCGAACGCGCUAACCAAUUGCGCCACGGAGGCCAUGU--- ((((.(((.............((((((.....)))))).((((.........))))....))).))))..--- ( -20.80, z-score = -1.50, R) >droSim1.chr2R 18623078 65 - 19596830 GUGGACUCGAACCACCAACCUUUCGGUUAACAGCCGAACGCGCUAACCAAUUGCGCCACGGAGGC-------- ((((.......))))...(((((((((.....)))))..((((.........))))....)))).-------- ( -18.60, z-score = -1.29, R) >droSec1.super_26 720597 70 + 826190 GUGGACUCGAACCACCAACCUUUCGGUUAACAGCCGAACGCGCUAACCAAUUGCGCCACGGAGGCCAUGU--- ((((.(((.............((((((.....)))))).((((.........))))....))).))))..--- ( -20.80, z-score = -1.50, R) >droYak2.chr2L 1925101 68 - 22324452 GUGGACUCGAACCACCAACCUUUCGGUUAACAGCCGAACGCGCUAACCAAUUGCGCCACGGAGGCCGU----- ..((.(((.............((((((.....)))))).((((.........))))....))).))..----- ( -18.80, z-score = -0.67, R) >droEre2.scaffold_4929 20292255 68 + 26641161 GUGGACUCGAACCACCAACCUUUCGGUUAACAGCCGAACGCGCUAACCAAUUGCGCCACGGAGGCUAU----- ((((.......))))...(((((((((.....)))))..((((.........))))....))))....----- ( -18.60, z-score = -1.32, R) >droAna3.scaffold_13266 1157561 65 - 19884421 GUGGACUCGAACCACCAACCUUUCGGUUAAAAGCCGAACGCGCUAACCGAUUGCGCCACGGAGGC-------- ((((.......))))...(((((((((.....)))))..((((.........))))....)))).-------- ( -18.60, z-score = -0.94, R) >dp4.chr3 18003717 68 + 19779522 GUGGACUCGAACCACCAACCUUUCGGUUAACAGCCGAACGCGCUAACCAAUUGCGCCACGGAGGCCAU----- ((((.(((.............((((((.....)))))).((((.........))))....))).))))----- ( -20.10, z-score = -1.82, R) >droPer1.super_73 214616 68 - 267513 GUGGACUCGAACCACCAACCUUUCGGUUAACAGCCGAACGCGCUAACCAAUUGCGCCACGGAGGCCAU----- ((((.(((.............((((((.....)))))).((((.........))))....))).))))----- ( -20.10, z-score = -1.82, R) >droWil1.scaffold_180700 773111 70 - 6630534 GUGGACUCGAACCACCAACCUUUCGGUUAACAGCCGAACGCGCUAACCGAUUGCGCCACGGAGGCGAUGU--- ((((.......))))....(..(((((((...((.....))..)))))))..)((((.....))))....--- ( -19.80, z-score = -0.60, R) >droVir3.scaffold_12875 4822565 70 + 20611582 GUGGACUCGAACCACCAACCUUUCGGUUAACAGCCGAACGCGCUAACCGAUUGCGCCACGGAGGCCAUGU--- ((((.(((.............((((((.....)))))).((((.........))))....))).))))..--- ( -20.80, z-score = -1.18, R) >droMoj3.scaffold_6496 1612958 67 - 26866924 GUGGACUCGAACCACCAACCUUUCGGUUAACAGCCGAACGCGCUAACCGAUUGCGCCACGGAGGCCA------ .(((.(((.............((((((.....)))))).((((.........))))....))).)))------ ( -18.90, z-score = -0.88, R) >droGri2.scaffold_15245 18152653 68 - 18325388 GUGGACUCGAACCACCAACCUUUCGGUUAACAGCCGAACGCGCUAACCAAUUGCGCCACGGAGGCUGU----- ((((.......)))).((((....)))).((((((....((((.........))))......))))))----- ( -21.60, z-score = -1.80, R) >anoGam1.chr2L 3862547 73 - 48795086 GUGGACUCGAACCACCAACCUUUCGGUUAACAGCCGAACGCGCUAACCGAUUGCGCCACGAAGGCGGUGAGAG .....(((..(((.((.....((((((.....)))))).((((.........))))......)).)))..))) ( -24.10, z-score = -1.88, R) >consensus GUGGACUCGAACCACCAACCUUUCGGUUAACAGCCGAACGCGCUAACCAAUUGCGCCACGGAGGCCAU_____ ((((.......))))...(((((((((.....)))))..((((.........))))....))))......... (-18.64 = -18.57 + -0.07)


| Location | 2,077,642 – 2,077,712 |
|---|---|
| Length | 70 |
| Sequences | 13 |
| Columns | 73 |
| Reading direction | reverse |
| Mean pairwise identity | 94.22 |
| Shannon entropy | 0.12272 |
| G+C content | 0.57979 |
| Mean single sequence MFE | -25.30 |
| Consensus MFE | -23.86 |
| Energy contribution | -23.86 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.52 |
| Structure conservation index | 0.94 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.93 |
| SVM RNA-class probability | 0.854773 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 2077642 70 - 21146708 ---ACAUGGCCUCCGUGGCGCAAUUGGUUAGCGCGUUCGGCUGUUAACCGAAAGGUUGGUGGUUCGAGUCCAC ---.(((((...)))))((((.........)))).((((((..((((((....))))))..).)))))..... ( -26.80, z-score = -1.81, R) >droSim1.chr2R 18623078 65 + 19596830 --------GCCUCCGUGGCGCAAUUGGUUAGCGCGUUCGGCUGUUAACCGAAAGGUUGGUGGUUCGAGUCCAC --------......(((((((.........)))).((((((..((((((....))))))..).)))))..))) ( -24.10, z-score = -1.56, R) >droSec1.super_26 720597 70 - 826190 ---ACAUGGCCUCCGUGGCGCAAUUGGUUAGCGCGUUCGGCUGUUAACCGAAAGGUUGGUGGUUCGAGUCCAC ---.(((((...)))))((((.........)))).((((((..((((((....))))))..).)))))..... ( -26.80, z-score = -1.81, R) >droYak2.chr2L 1925101 68 + 22324452 -----ACGGCCUCCGUGGCGCAAUUGGUUAGCGCGUUCGGCUGUUAACCGAAAGGUUGGUGGUUCGAGUCCAC -----..((.(((((..((((.........))))...))((..((((((....))))))..))..))).)).. ( -25.70, z-score = -1.37, R) >droEre2.scaffold_4929 20292255 68 - 26641161 -----AUAGCCUCCGUGGCGCAAUUGGUUAGCGCGUUCGGCUGUUAACCGAAAGGUUGGUGGUUCGAGUCCAC -----.........(((((((.........)))).((((((..((((((....))))))..).)))))..))) ( -24.10, z-score = -1.56, R) >droAna3.scaffold_13266 1157561 65 + 19884421 --------GCCUCCGUGGCGCAAUCGGUUAGCGCGUUCGGCUUUUAACCGAAAGGUUGGUGGUUCGAGUCCAC --------......(((((((.........)))).(((((((.((((((....)))))).)).)))))..))) ( -21.40, z-score = -0.64, R) >dp4.chr3 18003717 68 - 19779522 -----AUGGCCUCCGUGGCGCAAUUGGUUAGCGCGUUCGGCUGUUAACCGAAAGGUUGGUGGUUCGAGUCCAC -----.(((.(((((..((((.........))))...))((..((((((....))))))..))..))).))). ( -26.10, z-score = -1.89, R) >droPer1.super_73 214616 68 + 267513 -----AUGGCCUCCGUGGCGCAAUUGGUUAGCGCGUUCGGCUGUUAACCGAAAGGUUGGUGGUUCGAGUCCAC -----.(((.(((((..((((.........))))...))((..((((((....))))))..))..))).))). ( -26.10, z-score = -1.89, R) >droWil1.scaffold_180700 773111 70 + 6630534 ---ACAUCGCCUCCGUGGCGCAAUCGGUUAGCGCGUUCGGCUGUUAACCGAAAGGUUGGUGGUUCGAGUCCAC ---...........(((((((.........)))).((((((..((((((....))))))..).)))))..))) ( -24.10, z-score = -0.87, R) >droVir3.scaffold_12875 4822565 70 - 20611582 ---ACAUGGCCUCCGUGGCGCAAUCGGUUAGCGCGUUCGGCUGUUAACCGAAAGGUUGGUGGUUCGAGUCCAC ---.(((((...)))))((((.........)))).((((((..((((((....))))))..).)))))..... ( -26.80, z-score = -1.64, R) >droMoj3.scaffold_6496 1612958 67 + 26866924 ------UGGCCUCCGUGGCGCAAUCGGUUAGCGCGUUCGGCUGUUAACCGAAAGGUUGGUGGUUCGAGUCCAC ------(((.(((((..((((.........))))...))((..((((((....))))))..))..))).))). ( -25.80, z-score = -1.58, R) >droGri2.scaffold_15245 18152653 68 + 18325388 -----ACAGCCUCCGUGGCGCAAUUGGUUAGCGCGUUCGGCUGUUAACCGAAAGGUUGGUGGUUCGAGUCCAC -----((((((......((((.........))))....))))))(((((....)))))((((.......)))) ( -24.20, z-score = -1.29, R) >anoGam1.chr2L 3862547 73 + 48795086 CUCUCACCGCCUUCGUGGCGCAAUCGGUUAGCGCGUUCGGCUGUUAACCGAAAGGUUGGUGGUUCGAGUCCAC (((((((((((.....)))....(((((((((((.....)).)))))))))......)))))...)))..... ( -26.90, z-score = -1.78, R) >consensus _____AUGGCCUCCGUGGCGCAAUUGGUUAGCGCGUUCGGCUGUUAACCGAAAGGUUGGUGGUUCGAGUCCAC ..............(((((((.........)))).((((((..((((((....))))))..).)))))..))) (-23.86 = -23.86 + 0.00)



Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:03:17 2011