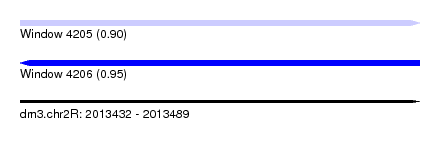

| Sequence ID | dm3.chr2R |
|---|---|
| Location | 2,013,432 – 2,013,489 |
| Length | 57 |
| Max. P | 0.950076 |

| Location | 2,013,432 – 2,013,489 |
|---|---|
| Length | 57 |
| Sequences | 5 |
| Columns | 57 |
| Reading direction | forward |
| Mean pairwise identity | 79.47 |
| Shannon entropy | 0.36646 |
| G+C content | 0.35170 |
| Mean single sequence MFE | -11.30 |
| Consensus MFE | -6.08 |
| Energy contribution | -6.56 |
| Covariance contribution | 0.48 |
| Combinations/Pair | 1.36 |
| Mean z-score | -2.34 |
| Structure conservation index | 0.54 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.13 |
| SVM RNA-class probability | 0.897073 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
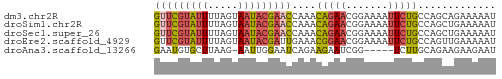
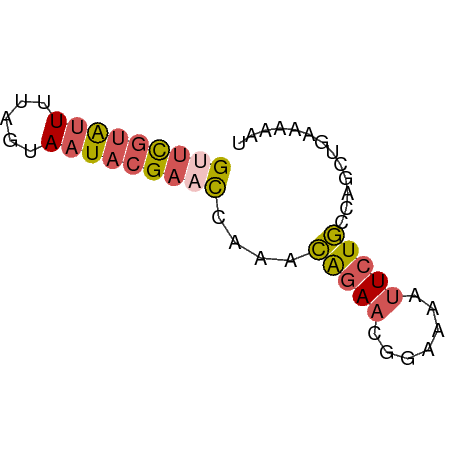
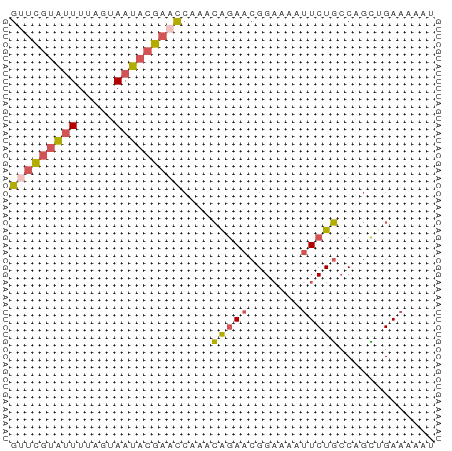
>dm3.chr2R 2013432 57 + 21146708 GUUCGUAUUUUAGUAAUACGAACCAAACAGAACGGAAAAUUCUGCCAGCAGAAAAAU (((((((((.....)))))))))................(((((....))))).... ( -14.30, z-score = -4.40, R) >droSim1.chr2R 910299 57 + 19596830 GUUCGUAUUUUAGUAAUACGAACCAAACAGAACGGAAAAUUCUGCCAGCUGAAAAAU (((((((((.....)))))))))....(((..((((....))))....)))...... ( -13.90, z-score = -3.73, R) >droSec1.super_26 656441 57 + 826190 GUUCGUAUUUUAGUAAUACGAACCAAACAGAACGGAAAAUUCUGCCAGCUGAAAAAU (((((((((.....)))))))))....(((..((((....))))....)))...... ( -13.90, z-score = -3.73, R) >droEre2.scaffold_4929 6320114 57 - 26641161 GUUCGUAUUUUAGUAAUACGAUUGAAACGGAACGGAAAAUUCUGCCAGUUGAAAAAU ..(((((((.....)))))))((..(((((..((((....)))))).)))..))... ( -9.40, z-score = -0.77, R) >droAna3.scaffold_13266 1100628 51 - 19884421 GAAUGUGCUUAAG-AAUUGGAAUCAGAAGAAUCGG-----UCUUGCAGAAGAAGAAU ...(.(((..(((-(....((.((....)).))..-----))))))).)........ ( -5.00, z-score = 0.93, R) >consensus GUUCGUAUUUUAGUAAUACGAACCAAACAGAACGGAAAAUUCUGCCAGCUGAAAAAU (((((((((.....)))))))))....(((((.......)))))............. ( -6.08 = -6.56 + 0.48)
| Location | 2,013,432 – 2,013,489 |
|---|---|
| Length | 57 |
| Sequences | 5 |
| Columns | 57 |
| Reading direction | reverse |
| Mean pairwise identity | 79.47 |
| Shannon entropy | 0.36646 |
| G+C content | 0.35170 |
| Mean single sequence MFE | -11.58 |
| Consensus MFE | -6.32 |
| Energy contribution | -8.32 |
| Covariance contribution | 2.00 |
| Combinations/Pair | 1.24 |
| Mean z-score | -2.53 |
| Structure conservation index | 0.55 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.56 |
| SVM RNA-class probability | 0.950076 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
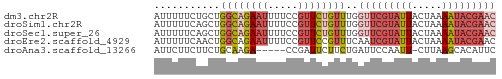
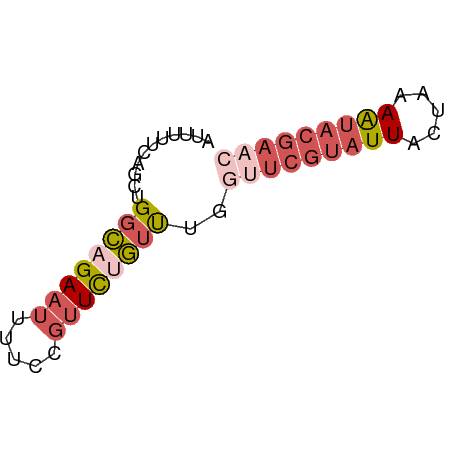
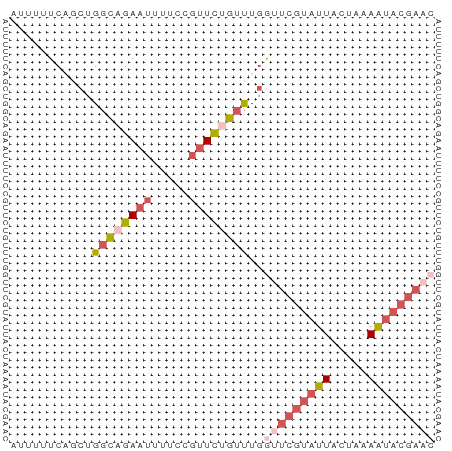
>dm3.chr2R 2013432 57 - 21146708 AUUUUUCUGCUGGCAGAAUUUUCCGUUCUGUUUGGUUCGUAUUACUAAAAUACGAAC .........(..(((((((.....)))))))..)(((((((((.....))))))))) ( -15.50, z-score = -3.95, R) >droSim1.chr2R 910299 57 - 19596830 AUUUUUCAGCUGGCAGAAUUUUCCGUUCUGUUUGGUUCGUAUUACUAAAAUACGAAC .........(..(((((((.....)))))))..)(((((((((.....))))))))) ( -15.50, z-score = -3.73, R) >droSec1.super_26 656441 57 - 826190 AUUUUUCAGCUGGCAGAAUUUUCCGUUCUGUUUGGUUCGUAUUACUAAAAUACGAAC .........(..(((((((.....)))))))..)(((((((((.....))))))))) ( -15.50, z-score = -3.73, R) >droEre2.scaffold_4929 6320114 57 + 26641161 AUUUUUCAACUGGCAGAAUUUUCCGUUCCGUUUCAAUCGUAUUACUAAAAUACGAAC ...........(((.((((.....)))).)))....(((((((.....))))))).. ( -7.40, z-score = -1.04, R) >droAna3.scaffold_13266 1100628 51 + 19884421 AUUCUUCUUCUGCAAGA-----CCGAUUCUUCUGAUUCCAAUU-CUUAAGCACAUUC ..........(((((((-----..((.((....)).))....)-)))..)))..... ( -4.00, z-score = -0.21, R) >consensus AUUUUUCAGCUGGCAGAAUUUUCCGUUCUGUUUGGUUCGUAUUACUAAAAUACGAAC ...........((((((((.....))))))))..(((((((((.....))))))))) ( -6.32 = -8.32 + 2.00)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:03:00 2011