
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 1,614,737 – 1,614,795 |
| Length | 58 |
| Max. P | 0.886236 |

| Location | 1,614,737 – 1,614,795 |
|---|---|
| Length | 58 |
| Sequences | 5 |
| Columns | 58 |
| Reading direction | forward |
| Mean pairwise identity | 75.84 |
| Shannon entropy | 0.39488 |
| G+C content | 0.56430 |
| Mean single sequence MFE | -16.88 |
| Consensus MFE | -9.50 |
| Energy contribution | -9.26 |
| Covariance contribution | -0.24 |
| Combinations/Pair | 1.42 |
| Mean z-score | -1.68 |
| Structure conservation index | 0.56 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.27 |
| SVM RNA-class probability | 0.619559 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
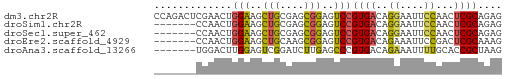
>dm3.chr2R 1614737 58 + 21146708 CCAGACUCGAACUGGAAGCUGCGAGCGGAGUCCGUGACAGGAAUUCCAACUCGCAGAG ((((.......))))...(((((((.(((((((......)).)))))..))))))).. ( -23.70, z-score = -3.02, R) >droSim1.chr2R 514702 51 + 19596830 -------CCAACUGGAAGCUGCGAGCGGAGUCCGUGACAGGAAUUCCAACUCGCAGAG -------((....))...(((((((.(((((((......)).)))))..))))))).. ( -20.50, z-score = -2.97, R) >droSec1.super_462 5792 51 - 10684 -------CCAACUGGAAGCUGCGAGCGGAGUCCGUGACAGGAAUUCCAACUCGCAGAG -------((....))...(((((((.(((((((......)).)))))..))))))).. ( -20.50, z-score = -2.97, R) >droEre2.scaffold_4929 5893671 51 - 26641161 -------CCAACUGGAAGCUGCAAGCGGAGUCCGUGACAGAAAUUCCGACUCGCAAAG -------.....(((((.(((...((((...))))..)))...))))).......... ( -12.00, z-score = -0.56, R) >droAna3.scaffold_13266 665816 51 - 19884421 -------UGGACUUGGAGUCGGAUCUUGAGCCCGUGACAGAAAUUUUGCACCGCUAAG -------..(((.....)))........(((..(((.((((...))))))).)))... ( -7.70, z-score = 1.12, R) >consensus _______CCAACUGGAAGCUGCGAGCGGAGUCCGUGACAGGAAUUCCAACUCGCAGAG .............(((..(((....)))..)))((((..((....))...)))).... ( -9.50 = -9.26 + -0.24)


| Location | 1,614,737 – 1,614,795 |
|---|---|
| Length | 58 |
| Sequences | 5 |
| Columns | 58 |
| Reading direction | reverse |
| Mean pairwise identity | 75.84 |
| Shannon entropy | 0.39488 |
| G+C content | 0.56430 |
| Mean single sequence MFE | -16.50 |
| Consensus MFE | -12.48 |
| Energy contribution | -12.20 |
| Covariance contribution | -0.28 |
| Combinations/Pair | 1.50 |
| Mean z-score | -1.43 |
| Structure conservation index | 0.76 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.07 |
| SVM RNA-class probability | 0.886236 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
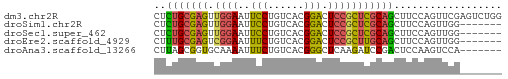
>dm3.chr2R 1614737 58 - 21146708 CUCUGCGAGUUGGAAUUCCUGUCACGGACUCCGCUCGCAGCUUCCAGUUCGAGUCUGG ..((((((((.(((..(((......))).)))))))))))...((((.......)))) ( -22.00, z-score = -2.19, R) >droSim1.chr2R 514702 51 - 19596830 CUCUGCGAGUUGGAAUUCCUGUCACGGACUCCGCUCGCAGCUUCCAGUUGG------- ..((((((((.(((..(((......))).)))))))))))...........------- ( -18.80, z-score = -2.24, R) >droSec1.super_462 5792 51 + 10684 CUCUGCGAGUUGGAAUUCCUGUCACGGACUCCGCUCGCAGCUUCCAGUUGG------- ..((((((((.(((..(((......))).)))))))))))...........------- ( -18.80, z-score = -2.24, R) >droEre2.scaffold_4929 5893671 51 + 26641161 CUUUGCGAGUCGGAAUUUCUGUCACGGACUCCGCUUGCAGCUUCCAGUUGG------- ..(((((((.((((...((((...)))).)))))))))))...........------- ( -14.90, z-score = -1.09, R) >droAna3.scaffold_13266 665816 51 + 19884421 CUUAGCGGUGCAAAAUUUCUGUCACGGGCUCAAGAUCCGACUCCAAGUCCA------- .......(((((.......)).)))(((((...((......))..))))).------- ( -8.00, z-score = 0.62, R) >consensus CUCUGCGAGUUGGAAUUCCUGUCACGGACUCCGCUCGCAGCUUCCAGUUGG_______ ..(((((((.((((..(((......))).))))))))))).................. (-12.48 = -12.20 + -0.28)
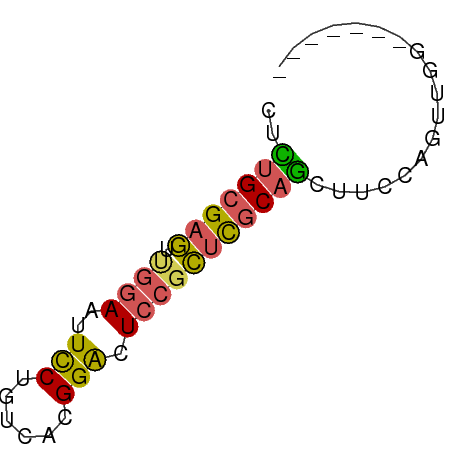

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:02:18 2011