| Sequence ID | dm3.chr2R |
|---|---|
| Location | 752,440 – 752,553 |
| Length | 113 |
| Max. P | 0.976344 |

| Location | 752,440 – 752,553 |
|---|---|
| Length | 113 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 59.89 |
| Shannon entropy | 0.56408 |
| G+C content | 0.54240 |
| Mean single sequence MFE | -45.03 |
| Consensus MFE | -17.47 |
| Energy contribution | -17.60 |
| Covariance contribution | 0.13 |
| Combinations/Pair | 1.41 |
| Mean z-score | -2.48 |
| Structure conservation index | 0.39 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.95 |
| SVM RNA-class probability | 0.976344 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 752440 113 - 21146708 CAUGCUGCUGAAAGGUGCCUUGAGAC------GUGCGCCUACGGUGAGGUUUGCUGGAGCGAACCUUCCGUAUGUG-CGUAGCUGUCGGUACCUUGGCAUCACGGUUAGUGAAGGAAUGA (((.(((((..((((((((....(((------((((((((((((.(((((((((....)))))))))))))).).)-))))..))))))))))))))).((((.....)))).)).))). ( -48.40, z-score = -3.69, R) >droAna3.scaffold_13230 1798603 113 - 3602488 AAUGCUGCUGAAAGGCAGAUUGUCCAG-----GUCCCUCGUU-GUCAGGCUCGACGGCUCCAGCAUAAGGUAUGCGAAACAGGUG-AAGUACCUGGGCGUGACCGUUGGAGAACGGUUGC ....(((((....)))))...((((((-----((.(.(((((-(((......))))))....(((((...))))).........)-).).))))))))(..((((((....))))))..) ( -44.70, z-score = -2.25, R) >droGri2.scaffold_15040 117870 119 - 162499 AAUGCUGCUUAAGGGCAGAUUGUCGCGUAGUCGACCCCCAUUGGUUAGGCUAAAUGGCGGCAGCCUGAGAUAUGUGUCGGAGGUG-AAAUACCUCGGCAUUGUGUUUGGCGAGAGGAUGU ....(((((....)))))..((((((.(((((((((......)))).)))))....))))))(((.((.(((.((((((.(((((-...)))))))))))))).)).))).......... ( -42.00, z-score = -1.49, R) >consensus AAUGCUGCUGAAAGGCAGAUUGUCACG_____GUCCCCCAUUGGUCAGGCUAGAUGGCGCCAGCCUAAGGUAUGUG_CGCAGGUG_AAGUACCUCGGCAUCACGGUUGGAGAAAGGAUGA ....(((((....)))))..........................(((((((.((....)).)))))))(((((((....((((((....)))))).)))).)))................ (-17.47 = -17.60 + 0.13)

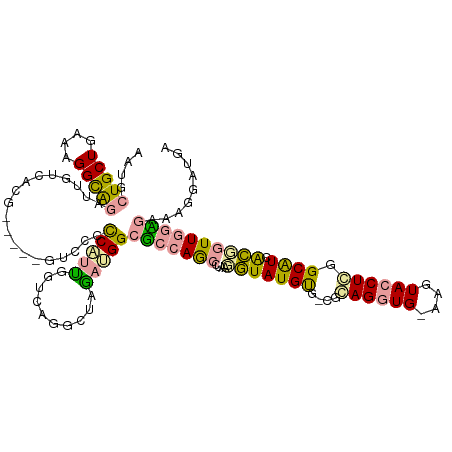
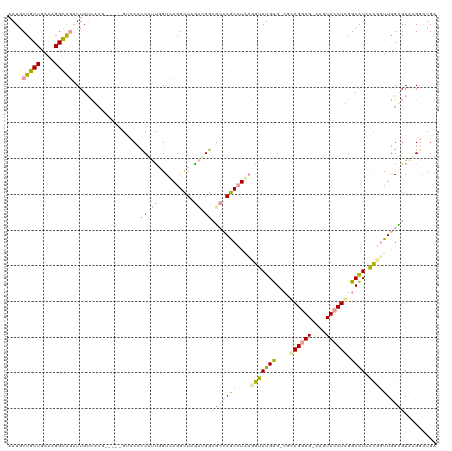
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:01:31 2011