
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 642,814 – 642,867 |
| Length | 53 |
| Max. P | 0.937346 |

| Location | 642,814 – 642,867 |
|---|---|
| Length | 53 |
| Sequences | 4 |
| Columns | 54 |
| Reading direction | forward |
| Mean pairwise identity | 53.89 |
| Shannon entropy | 0.74823 |
| G+C content | 0.32545 |
| Mean single sequence MFE | -8.55 |
| Consensus MFE | -2.12 |
| Energy contribution | -2.12 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.40 |
| Mean z-score | -1.90 |
| Structure conservation index | 0.25 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.58 |
| SVM RNA-class probability | 0.749269 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
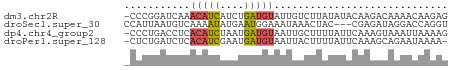
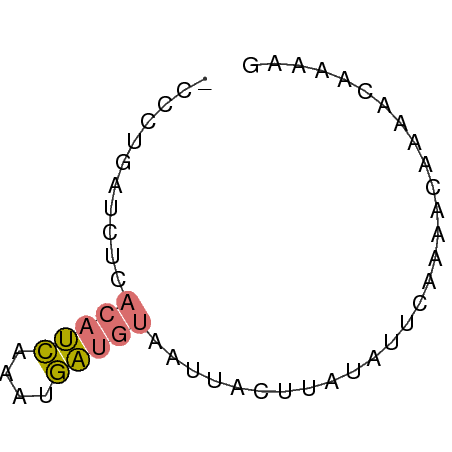
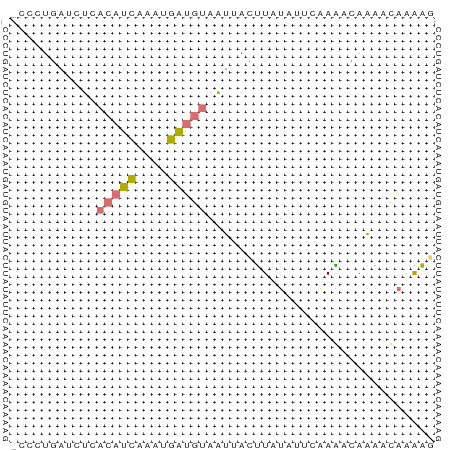
>dm3.chr2R 642814 53 + 21146708 -CCCGGAUCAAACAUCAUCUGAUGUAUUGUCUUAUAUACAAGACAAAACAAGAG -..(((((........))))).(((.(((((((......))))))).))).... ( -11.60, z-score = -2.89, R) >droSec1.super_30 421996 51 - 664411 CCAUUAAUGUCAAAAUAUGAAUGGAAAUAAACUAC---CGAGAUAGGACCAGGU (((((.((((....)))).)))))..........(---(......))....... ( -6.60, z-score = -0.36, R) >dp4.chr4_group2 150791 53 - 1235136 -CCCUGACCUCACAUCUAAUGAUGUAAUUGCUUUUAUUCAAAGUAAAUUAAAAG -..........(((((....)))))..(((((((.....)))))))........ ( -8.10, z-score = -2.83, R) >droPer1.super_128 26025 52 + 112355 -CUCUGAUCUCACAUCGAAUGAUGUAAUUACUUUUAUUCAAAGCAGAAUAAAA- -..........(((((....))))).......(((((((......))))))).- ( -7.90, z-score = -1.52, R) >consensus _CCCUGAUCUCACAUCAAAUGAUGUAAUUACUUAUAUUCAAAACAAAACAAAAG ...........(((((....)))))............................. ( -2.12 = -2.12 + 0.00)
| Location | 642,814 – 642,867 |
|---|---|
| Length | 53 |
| Sequences | 4 |
| Columns | 54 |
| Reading direction | reverse |
| Mean pairwise identity | 53.89 |
| Shannon entropy | 0.74823 |
| G+C content | 0.32545 |
| Mean single sequence MFE | -9.48 |
| Consensus MFE | -4.03 |
| Energy contribution | -3.90 |
| Covariance contribution | -0.13 |
| Combinations/Pair | 1.43 |
| Mean z-score | -1.27 |
| Structure conservation index | 0.42 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.41 |
| SVM RNA-class probability | 0.937346 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
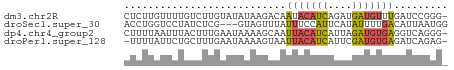
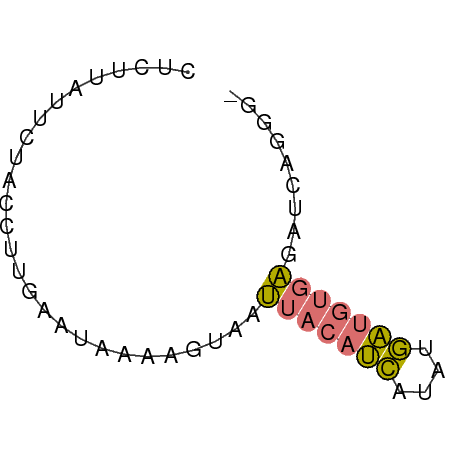
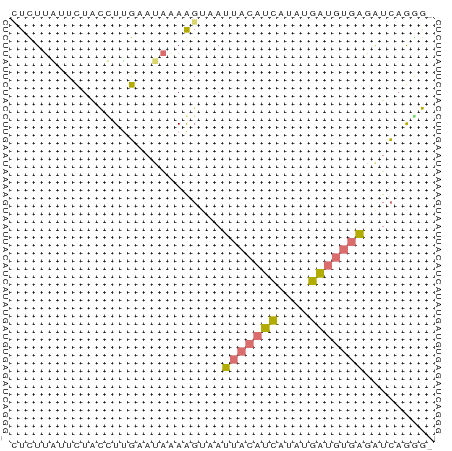
>dm3.chr2R 642814 53 - 21146708 CUCUUGUUUUGUCUUGUAUAUAAGACAAUACAUCAGAUGAUGUUUGAUCCGGG- (((.(((.((((((((....)))))))).)))((((((...))))))...)))- ( -11.90, z-score = -1.56, R) >droSec1.super_30 421996 51 + 664411 ACCUGGUCCUAUCUCG---GUAGUUUAUUUCCAUUCAUAUUUUGACAUUAAUGG (((.((......)).)---)).........(((((.((........)).))))) ( -4.40, z-score = 0.83, R) >dp4.chr4_group2 150791 53 + 1235136 CUUUUAAUUUACUUUGAAUAAAAGCAAUUACAUCAUUAGAUGUGAGGUCAGGG- ...........((((((..........(((((((....)))))))..))))))- ( -9.30, z-score = -1.74, R) >droPer1.super_128 26025 52 - 112355 -UUUUAUUCUGCUUUGAAUAAAAGUAAUUACAUCAUUCGAUGUGAGAUCAGAG- -((((((((......))))))))(...(((((((....)))))))...)....- ( -12.30, z-score = -2.60, R) >consensus CUCUUAUUCUACCUUGAAUAAAAGUAAUUACAUCAUAUGAUGUGAGAUCAGGG_ ...........................(((((((....)))))))......... ( -4.03 = -3.90 + -0.13)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:01:22 2011