
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 2,125,638 – 2,125,739 |
| Length | 101 |
| Max. P | 0.959917 |

| Location | 2,125,638 – 2,125,739 |
|---|---|
| Length | 101 |
| Sequences | 6 |
| Columns | 103 |
| Reading direction | forward |
| Mean pairwise identity | 77.25 |
| Shannon entropy | 0.44233 |
| G+C content | 0.32928 |
| Mean single sequence MFE | -19.32 |
| Consensus MFE | -10.63 |
| Energy contribution | -11.69 |
| Covariance contribution | 1.06 |
| Combinations/Pair | 1.35 |
| Mean z-score | -2.20 |
| Structure conservation index | 0.55 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.67 |
| SVM RNA-class probability | 0.959917 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
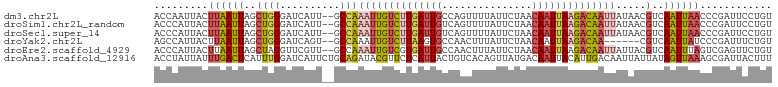
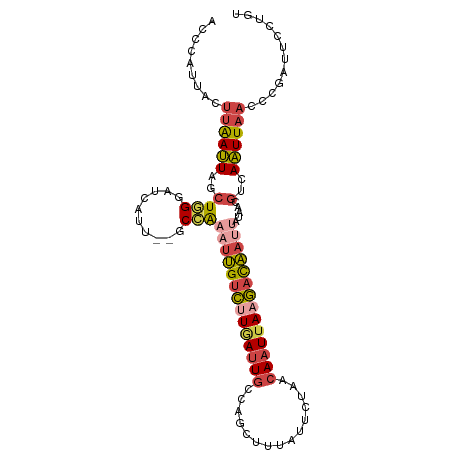
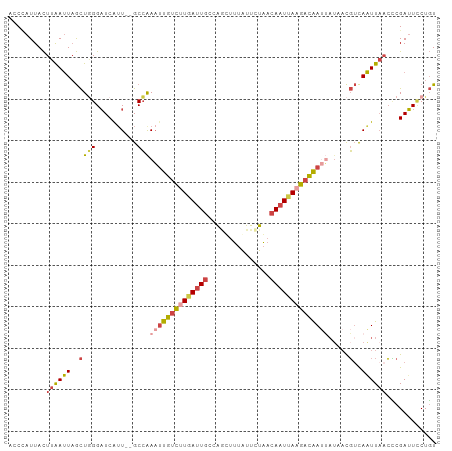
>dm3.chr2L 2125638 101 + 23011544 ACCAAUUACUUAAUUAGCUGGGAUCAUU--GCCAAAUUGUCUUGAUUGCCAGUUUUAUUCUAACAAUUAAGACAAUUAUAACGUCAAUUAACCCGAUUCCUGU ................((.((((((...--....((((((((((((((..((.......))..)))))))))))))).....((......))..)))))).)) ( -22.30, z-score = -3.46, R) >droSim1.chr2L_random 111125 101 + 909653 ACCCAUUACUUAAUUAGCUGGGAUCAUU--GCCAAAUUGUCUUGAUUGUCAGUUUUAUUCUAACAAUUAAGACAAUUAUAACGUCAAUUAACCCGAUUCCUGU ................((.((((((...--....(((((((((((((((.((.......)).))))))))))))))).....((......))..)))))).)) ( -24.70, z-score = -4.29, R) >droSec1.super_14 2055148 101 + 2068291 ACCCAUUACUUAAUUAGCUGGGAUCAUU--GCCAAAUUGUCUUGAUUGUCAGUUUUAUUCUAACAAUUAAGACAAUUAUAACGUCAAUUAACCCGAUUCCUGU ................((.((((((...--....(((((((((((((((.((.......)).))))))))))))))).....((......))..)))))).)) ( -24.70, z-score = -4.29, R) >droYak2.chr2L 2097031 95 + 22324452 AGCCAUUACUUAAUUAGCUGGGAUCAGU--GCCAAAUUGUCUUAAGUGCCAACUUUAUUCUAACAAUUAAGACAA------CGUCAAUUAUCCCGAUUUCUGU ..................((((((.(((--......(((((((((.((...............)).)))))))))------.....)))))))))........ ( -15.36, z-score = -0.68, R) >droEre2.scaffold_4929 2160038 101 + 26641161 ACCCAUUACUUAAUUAGCUAGGUUCGUU--GCCAAAUUGUCGUGAUUGCCAACUUUAUUCUAACAAUUAAGACAAUUAUUACGUCAAUUUAGUCGAGUUCUGU ...((..(((..(((((...(((.....--))).(((((((.((((((...............)))))).)))))))...........)))))..)))..)). ( -18.16, z-score = -1.57, R) >droAna3.scaffold_12916 9832715 103 - 16180835 ACCUAUUAUUUGACUCAUUUGGAUCAUUCUGCAGAUACGUUCUCAUUACUGUCACAGUUAUGACAAUUACAUUGACAAUUAUUAUAGUUAAAGCGAUUACUUU ........(((((((.(((((.(......).)))))..(((.((((.((((...)))).)))).)))..................)))))))........... ( -10.70, z-score = 1.08, R) >consensus ACCCAUUACUUAAUUAGCUGGGAUCAUU__GCCAAAUUGUCUUGAUUGCCAGCUUUAUUCUAACAAUUAAGACAAUUAUAACGUCAAUUAACCCGAUUCCUGU ................((.((((((.........((((((((((((((...............)))))))))))))).....(........)..)))))).)) (-10.63 = -11.69 + 1.06)
| Location | 2,125,638 – 2,125,739 |
|---|---|
| Length | 101 |
| Sequences | 6 |
| Columns | 103 |
| Reading direction | reverse |
| Mean pairwise identity | 77.25 |
| Shannon entropy | 0.44233 |
| G+C content | 0.32928 |
| Mean single sequence MFE | -22.97 |
| Consensus MFE | -11.39 |
| Energy contribution | -12.20 |
| Covariance contribution | 0.81 |
| Combinations/Pair | 1.38 |
| Mean z-score | -2.14 |
| Structure conservation index | 0.50 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.18 |
| SVM RNA-class probability | 0.905487 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
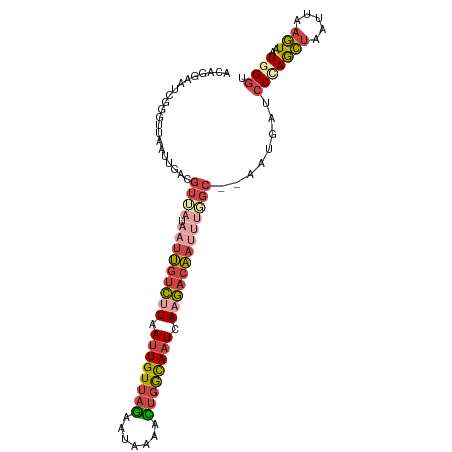
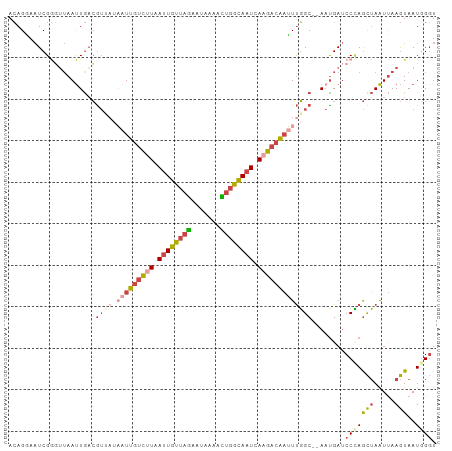
>dm3.chr2L 2125638 101 - 23011544 ACAGGAAUCGGGUUAAUUGACGUUAUAAUUGUCUUAAUUGUUAGAAUAAAACUGGCAAUCAAGACAAUUUGGC--AAUGAUCCCAGCUAAUUAAGUAAUUGGU .........(((...((((.......(((((((((.((((((((.......)))))))).)))))))))...)--)))...))).(((((((....))))))) ( -27.90, z-score = -3.67, R) >droSim1.chr2L_random 111125 101 - 909653 ACAGGAAUCGGGUUAAUUGACGUUAUAAUUGUCUUAAUUGUUAGAAUAAAACUGACAAUCAAGACAAUUUGGC--AAUGAUCCCAGCUAAUUAAGUAAUGGGU ......((((.(((....)))((((.(((((((((.((((((((.......)))))))).)))))))))))))--..))))(((((((.....)))..)))). ( -25.90, z-score = -3.04, R) >droSec1.super_14 2055148 101 - 2068291 ACAGGAAUCGGGUUAAUUGACGUUAUAAUUGUCUUAAUUGUUAGAAUAAAACUGACAAUCAAGACAAUUUGGC--AAUGAUCCCAGCUAAUUAAGUAAUGGGU ......((((.(((....)))((((.(((((((((.((((((((.......)))))))).)))))))))))))--..))))(((((((.....)))..)))). ( -25.90, z-score = -3.04, R) >droYak2.chr2L 2097031 95 - 22324452 ACAGAAAUCGGGAUAAUUGACG------UUGUCUUAAUUGUUAGAAUAAAGUUGGCACUUAAGACAAUUUGGC--ACUGAUCCCAGCUAAUUAAGUAAUGGCU .........(((((...((..(------(((((((((.((((((.......)))))).))))))))))....)--)...)))))(((((.........))))) ( -25.20, z-score = -2.73, R) >droEre2.scaffold_4929 2160038 101 - 26641161 ACAGAACUCGACUAAAUUGACGUAAUAAUUGUCUUAAUUGUUAGAAUAAAGUUGGCAAUCACGACAAUUUGGC--AACGAACCUAGCUAAUUAAGUAAUGGGU .....(((((........((((.......))))(((((((((((......((((.......))))..((((..--..)))).))))).))))))....))))) ( -19.00, z-score = -0.98, R) >droAna3.scaffold_12916 9832715 103 + 16180835 AAAGUAAUCGCUUUAACUAUAAUAAUUGUCAAUGUAAUUGUCAUAACUGUGACAGUAAUGAGAACGUAUCUGCAGAAUGAUCCAAAUGAGUCAAAUAAUAGGU (((((....)))))..((((..((.(((.(......((((((((....))))))))..((.((.(((.((....))))).)))).....).))).)))))).. ( -13.90, z-score = 0.60, R) >consensus ACAGGAAUCGGGUUAAUUGACGUUAUAAUUGUCUUAAUUGUUAGAAUAAAACUGGCAAUCAAGACAAUUUGGC__AAUGAUCCCAGCUAAUUAAGUAAUGGGU ..........................(((((((((.((((((((.......)))))))).)))))))))............(((((((.....)))..)))). (-11.39 = -12.20 + 0.81)


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:10:15 2011