
| Sequence ID | dm3.chr2RHet |
|---|---|
| Location | 2,583,195 – 2,583,251 |
| Length | 56 |
| Max. P | 0.989856 |

| Location | 2,583,195 – 2,583,251 |
|---|---|
| Length | 56 |
| Sequences | 7 |
| Columns | 58 |
| Reading direction | forward |
| Mean pairwise identity | 64.07 |
| Shannon entropy | 0.70680 |
| G+C content | 0.65175 |
| Mean single sequence MFE | -22.69 |
| Consensus MFE | -14.78 |
| Energy contribution | -15.09 |
| Covariance contribution | 0.31 |
| Combinations/Pair | 1.38 |
| Mean z-score | -1.03 |
| Structure conservation index | 0.65 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.39 |
| SVM RNA-class probability | 0.989856 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
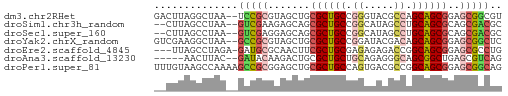
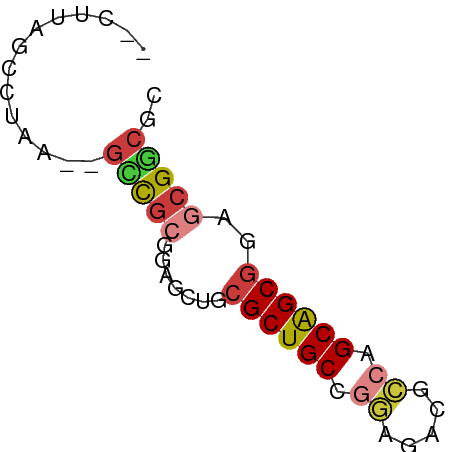
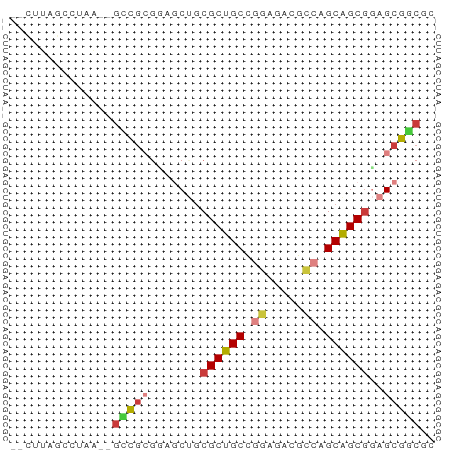
>dm3.chr2RHet 2583195 56 + 3288761 GACUUAGGCUAA--UCCGCGUAGCUGCGCUGCCGGGUACGCCAGCAGCGGAGCGGCGU ......((....--.))((((.(((.((((((.((.....)).)))))).))).)))) ( -24.80, z-score = -0.54, R) >droSim1.chr3h_random 107942 54 - 1452968 --CUUAGCCUAA--GUCGAAGAGCAGCGCUGCCGGCAUAGCCUGCAGCGCAGCGACGC --..........--((((.......(((((((.(((...))).)))))))..)))).. ( -22.50, z-score = -1.28, R) >droSec1.super_160 33429 54 - 40611 --CUUAGCCUAA--GUCGAGGAGCAGCGCUGCCGGCAUAGCCUGCAGCGCAGCGACGC --.....(((..--....))).((.(((((((.(((...))).))))))).))..... ( -23.50, z-score = -1.19, R) >droYak2.chrX_random 539170 56 + 1802292 GUCGAAGGCUAA--GCCGCGUAGCUGCGCUGCCGGAUACGACAGCAGCGGAGCGGCUC ...........(--(((((....((((((((.((....)).)))).)))).)))))). ( -25.30, z-score = -1.23, R) >droEre2.scaffold_4845 338015 54 + 22589142 ---UUAGCCUAGA-GAUGCGCAACUUCGCUGCGAGAGAGACCGGCAGCGGAGCGCCUG ---..........-...((((...((((((((..(......).))))))))))))... ( -17.80, z-score = -0.45, R) >droAna3.scaffold_13230 2044947 51 - 3602488 -----AACUUAC--GAUACAAGACUGCGCUGCUGCAGAGGGCAGCGGCUGAGCGUCAG -----.......--.......(((.((((((((((.....))))))))...))))).. ( -19.00, z-score = -1.55, R) >droPer1.super_81 267500 58 - 277874 UUUGUAAGCCAAAAGCCGCGGAGCUGCGCUGCCAGUGACGCCGGCAGCGGAGCGGCAG .......(((.....(....).(((.(((((((.(.....).))))))).)))))).. ( -25.90, z-score = -0.96, R) >consensus __CUUAGCCUAA__GCCGCGGAGCUGCGCUGCCGGAGACGCCAGCAGCGGAGCGGCGC ..............(((((.......((((((.((.....)).))))))..))))).. (-14.78 = -15.09 + 0.31)
| Location | 2,583,195 – 2,583,251 |
|---|---|
| Length | 56 |
| Sequences | 7 |
| Columns | 58 |
| Reading direction | reverse |
| Mean pairwise identity | 64.07 |
| Shannon entropy | 0.70680 |
| G+C content | 0.65175 |
| Mean single sequence MFE | -21.80 |
| Consensus MFE | -14.42 |
| Energy contribution | -14.53 |
| Covariance contribution | 0.11 |
| Combinations/Pair | 1.27 |
| Mean z-score | -0.87 |
| Structure conservation index | 0.66 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.15 |
| SVM RNA-class probability | 0.983916 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
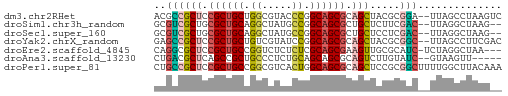
>dm3.chr2RHet 2583195 56 - 3288761 ACGCCGCUCCGCUGCUGGCGUACCCGGCAGCGCAGCUACGCGGA--UUAGCCUAAGUC .(((.(((.(((((((((.....))))))))).)))...)))..--............ ( -24.60, z-score = -1.39, R) >droSim1.chr3h_random 107942 54 + 1452968 GCGUCGCUGCGCUGCAGGCUAUGCCGGCAGCGCUGCUCUUCGAC--UUAGGCUAAG-- ..(((((.(((((((.(((...))).))))))).)).....)))--..........-- ( -22.50, z-score = -0.50, R) >droSec1.super_160 33429 54 + 40611 GCGUCGCUGCGCUGCAGGCUAUGCCGGCAGCGCUGCUCCUCGAC--UUAGGCUAAG-- ..(((((.(((((((.(((...))).))))))).)).....)))--..........-- ( -22.50, z-score = -0.35, R) >droYak2.chrX_random 539170 56 - 1802292 GAGCCGCUCCGCUGCUGUCGUAUCCGGCAGCGCAGCUACGCGGC--UUAGCCUUCGAC (((((((...((((((((((....)))))..)))))...)))))--)).......... ( -25.70, z-score = -2.09, R) >droEre2.scaffold_4845 338015 54 - 22589142 CAGGCGCUCCGCUGCCGGUCUCUCUCGCAGCGAAGUUGCGCAUC-UCUAGGCUAA--- ...((((..((((((.((.....)).)))))).....))))...-..........--- ( -18.20, z-score = -0.50, R) >droAna3.scaffold_13230 2044947 51 + 3602488 CUGACGCUCAGCCGCUGCCCUCUGCAGCAGCGCAGUCUUGUAUC--GUAAGUU----- (((.((((.....(((((.....)))))))))))).........--.......----- ( -14.00, z-score = -0.15, R) >droPer1.super_81 267500 58 + 277874 CUGCCGCUCCGCUGCCGGCGUCACUGGCAGCGCAGCUCCGCGGCUUUUGGCUUACAAA ((((.(((.(((((((((.....))))))))).)))...))))..((((.....)))) ( -25.10, z-score = -1.12, R) >consensus CCGCCGCUCCGCUGCUGGCGUCUCCGGCAGCGCAGCUCCGCGGC__UUAGGCUAAG__ .....(((.((((((.((.....)).)))))).)))...................... (-14.42 = -14.53 + 0.11)


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:00:34 2011