
| Sequence ID | dm3.chr2RHet |
|---|---|
| Location | 2,079,964 – 2,080,020 |
| Length | 56 |
| Max. P | 0.998735 |

| Location | 2,079,964 – 2,080,020 |
|---|---|
| Length | 56 |
| Sequences | 3 |
| Columns | 62 |
| Reading direction | forward |
| Mean pairwise identity | 74.73 |
| Shannon entropy | 0.34735 |
| G+C content | 0.39354 |
| Mean single sequence MFE | -16.57 |
| Consensus MFE | -15.30 |
| Energy contribution | -15.30 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -3.36 |
| Structure conservation index | 0.92 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 3.47 |
| SVM RNA-class probability | 0.998735 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
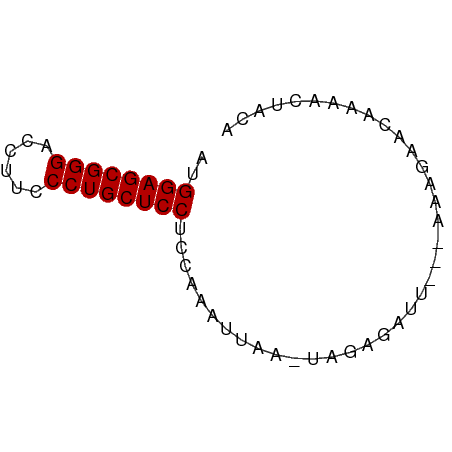
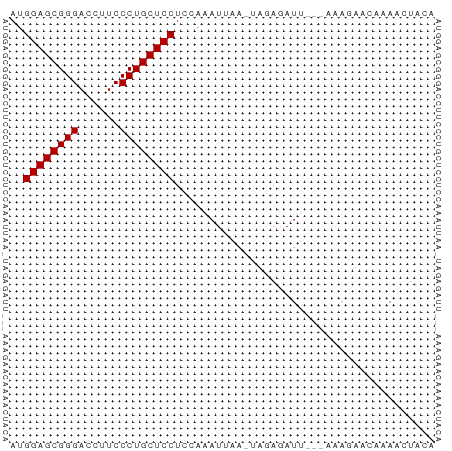
>dm3.chr2RHet 2079964 56 + 3288761 AUGGAGCGGGACCUUCCCUGCUCCUC-AAAUUGA-UUGAAAU----AAAGAACAUAAAUACA ..((((((((......))))))))((-((.....-))))...----................ ( -15.80, z-score = -3.56, R) >droSim1.chr3h_random 901487 62 + 1452968 AUGGAGCGGGACCUUCCCUGCUCCUCCAAAUUAAAUCGAGAUUUUAAAGGGAUAAAACUACA ..((((((((......))))))))(((...(((((.......)))))..))).......... ( -18.50, z-score = -2.60, R) >droSec1.super_174 3855 57 - 34763 AUGGAGCGGGACCUUCCCUGCUCCUACAAAUAAA-UAAAGAUU----GAAUUCAAAACUAAA ..((((((((......))))))))..........-......((----(....)))....... ( -15.40, z-score = -3.90, R) >consensus AUGGAGCGGGACCUUCCCUGCUCCUCCAAAUUAA_UAGAGAUU___AAAGAACAAAACUACA ..((((((((......))))))))...................................... (-15.30 = -15.30 + -0.00)

| Location | 2,079,964 – 2,080,020 |
|---|---|
| Length | 56 |
| Sequences | 3 |
| Columns | 62 |
| Reading direction | reverse |
| Mean pairwise identity | 74.73 |
| Shannon entropy | 0.34735 |
| G+C content | 0.39354 |
| Mean single sequence MFE | -16.07 |
| Consensus MFE | -14.80 |
| Energy contribution | -14.80 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.07 |
| Structure conservation index | 0.92 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.38 |
| SVM RNA-class probability | 0.989607 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
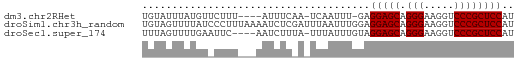
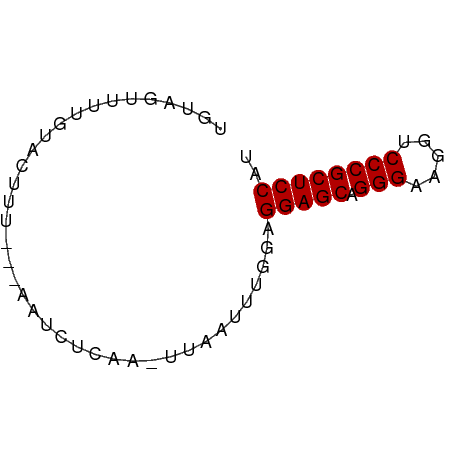
>dm3.chr2RHet 2079964 56 - 3288761 UGUAUUUAUGUUCUUU----AUUUCAA-UCAAUUU-GAGGAGCAGGGAAGGUCCCGCUCCAU ................----...((((-.....))-))(((((.(((.....)))))))).. ( -16.20, z-score = -2.55, R) >droSim1.chr3h_random 901487 62 - 1452968 UGUAGUUUUAUCCCUUUAAAAUCUCGAUUUAAUUUGGAGGAGCAGGGAAGGUCCCGCUCCAU ..........(((..(((((.......)))))...)))(((((.(((.....)))))))).. ( -17.20, z-score = -1.45, R) >droSec1.super_174 3855 57 + 34763 UUUAGUUUUGAAUUC----AAUCUUUA-UUUAUUUGUAGGAGCAGGGAAGGUCCCGCUCCAU ...............----........-..........(((((.(((.....)))))))).. ( -14.80, z-score = -2.20, R) >consensus UGUAGUUUUGUACUUU___AAUCUCAA_UUAAUUUGGAGGAGCAGGGAAGGUCCCGCUCCAU ......................................(((((.(((.....)))))))).. (-14.80 = -14.80 + -0.00)
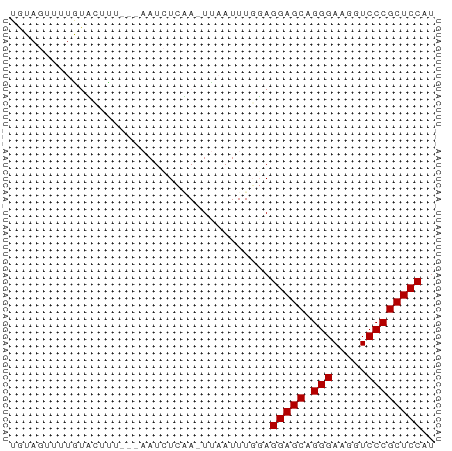
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:00:17 2011