
| Sequence ID | dm3.chr2RHet |
|---|---|
| Location | 1,986,601 – 1,986,679 |
| Length | 78 |
| Max. P | 0.864136 |

| Location | 1,986,601 – 1,986,679 |
|---|---|
| Length | 78 |
| Sequences | 4 |
| Columns | 78 |
| Reading direction | reverse |
| Mean pairwise identity | 62.82 |
| Shannon entropy | 0.61404 |
| G+C content | 0.35456 |
| Mean single sequence MFE | -14.69 |
| Consensus MFE | -8.76 |
| Energy contribution | -7.51 |
| Covariance contribution | -1.25 |
| Combinations/Pair | 1.65 |
| Mean z-score | -0.79 |
| Structure conservation index | 0.60 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.97 |
| SVM RNA-class probability | 0.864136 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
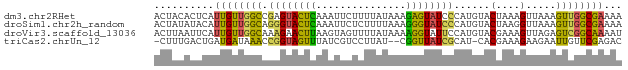
>dm3.chr2RHet 1986601 78 - 3288761 ACUACACUCAUUGUUGGCCGAGUACUCAAAUUCUUUUAUAAAGAGUAUCCCAUGUACUAAAGUUAAAGUUGGCGAAAA ..........((((..(((.(((((....((((((.....)))))).......)))))...).....))..))))... ( -11.80, z-score = -0.12, R) >droSim1.chr2h_random 502149 78 - 3178526 ACUAUAUACAUUGUUGGCAGGGUACUCAAAUUCUCUUUUAAAGGGUAUCCCAUGUACUAAGGUUAAAGUUGGCGAAAA ..........((((..((.((((((((...............)))))))).....((....))....))..))))... ( -15.56, z-score = -0.52, R) >droVir3.scaffold_13036 606669 78 - 1135500 ACUUAAUUCAUUGUUGGCAAAGAACUUAAGUAGUUUUAUAAAAGGUAUUCCAUGUACGAAAGUUAGAGUCGGCAAAAU ..........((((((((..((((((.....))))))......((....))....((....))....))))))))... ( -14.40, z-score = -1.62, R) >triCas2.chrUn_12 355620 74 - 931581 -CUUUGACUGAUGAUAAACCGGUAGUUUAUCGUCCUUAU--CGGUUAUCGCAU-CACGAAAGAAGAAUUGUUCGAGAC -(((.((((((((...(((((((((..........))))--)))))....)))-))(....).......))).))).. ( -17.00, z-score = -0.88, R) >consensus ACUAUAUUCAUUGUUGGCAGAGUACUCAAAUUCUCUUAUAAAGGGUAUCCCAUGUACGAAAGUUAAAGUUGGCGAAAA ..........((((((((.((((((((...............))))))))......(....).....))))))))... ( -8.76 = -7.51 + -1.25)


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:00:15 2011