| Sequence ID | dm3.chr2RHet |
|---|---|
| Location | 1,552,681 – 1,552,750 |
| Length | 69 |
| Max. P | 0.948156 |

| Location | 1,552,681 – 1,552,750 |
|---|---|
| Length | 69 |
| Sequences | 3 |
| Columns | 80 |
| Reading direction | forward |
| Mean pairwise identity | 56.17 |
| Shannon entropy | 0.61969 |
| G+C content | 0.35947 |
| Mean single sequence MFE | -14.37 |
| Consensus MFE | -7.50 |
| Energy contribution | -7.07 |
| Covariance contribution | -0.43 |
| Combinations/Pair | 1.54 |
| Mean z-score | -1.36 |
| Structure conservation index | 0.52 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.54 |
| SVM RNA-class probability | 0.948156 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2RHet 1552681 69 + 3288761 CUUAAUAUGAGGUUUAUUGAUCUCAAAAAAGCGGGAAAUAACAAGCUUGU--UAAGAAUAUUCUUGUCUUG--------- ((((((.(((((((....)))))))...((((..(......)..))))))--))))...............--------- ( -13.40, z-score = -1.85, R) >droGri2.scaffold_15116 579334 78 + 1808639 CUGAGCAUGAGAUUUGUUGAUCUCAAGACGGCUGAGAACAAGAAUCUGGU--GAAAAAUAUAAUUGUCGUCUGAAGACAC .....(((.((((((.(((.(((((.......))))).))))))))).))--)...........((((.......)))). ( -18.50, z-score = -1.45, R) >droVir3.scaffold_13324 720590 77 + 2960039 CUAAAUAUGUGUUUCUUAGACCUCAAACCGGCUUCCAACAAUAAGGAAGCACUAAAAAUAAACCUUUGUGCCGUCAA--- ..................(((.........((((((........)))))).......((((....))))...)))..--- ( -11.20, z-score = -0.78, R) >consensus CUAAAUAUGAGAUUUAUUGAUCUCAAAACGGCUGAAAACAACAAGCUAGU__UAAAAAUAUACUUGUCUUC_G___A___ .......(((((((....))))))).....((((((........)))))).............................. ( -7.50 = -7.07 + -0.43)

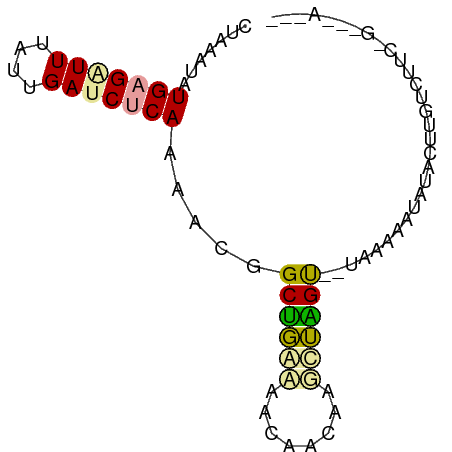
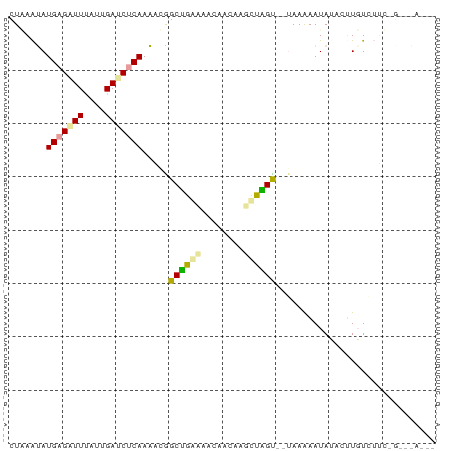
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:59:50 2011