
| Sequence ID | dm3.chr2RHet |
|---|---|
| Location | 891,067 – 891,225 |
| Length | 158 |
| Max. P | 0.800262 |

| Location | 891,067 – 891,155 |
|---|---|
| Length | 88 |
| Sequences | 4 |
| Columns | 88 |
| Reading direction | forward |
| Mean pairwise identity | 85.34 |
| Shannon entropy | 0.21569 |
| G+C content | 0.54230 |
| Mean single sequence MFE | -24.93 |
| Consensus MFE | -22.49 |
| Energy contribution | -22.49 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.37 |
| Structure conservation index | 0.90 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.73 |
| SVM RNA-class probability | 0.800262 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
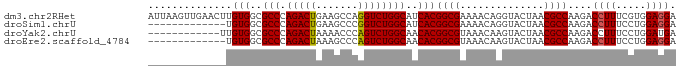
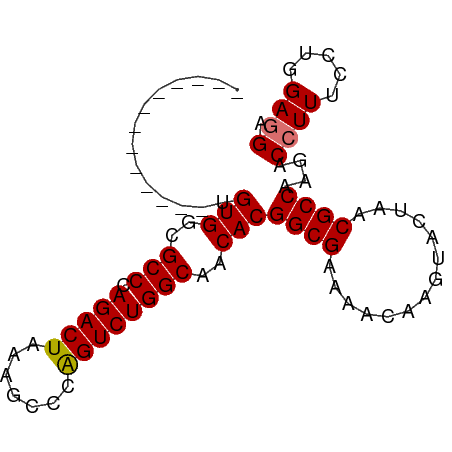
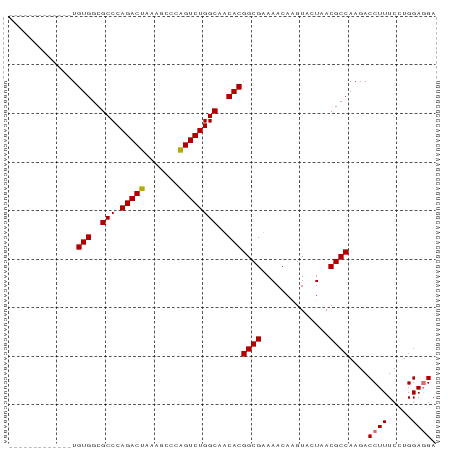
>dm3.chr2RHet 891067 88 + 3288761 AUUAAGUUGAACUUGUGGCGCCCAGACUGAAGCCAGGUCUGGCAUCACGGCGAAAACAGGUACUAACGCCAAGACCUUUCGUGGAGGA .....((((.((((((..((((.....(((.((((....)))).))).))))...))))))..)))).......((((.....)))). ( -26.20, z-score = -0.59, R) >droSim1.chrU 4942486 75 - 15797150 -------------UGUGGCGCCCAGACUGAAGCCCGGUCUGGCAUCACGGCGAAAACAGGUACUAACGCCAAGACCUUUCCUGGAGGA -------------.(((..(((.((((((.....)))))))))..)))((((..............))))....((((.....)))). ( -26.34, z-score = -1.18, R) >droYak2.chrU 2414593 76 + 28119190 ------------UUGUGGCGCCCAGACUAAAACCCAGUCUGGCAACACGGCGUAAACAAGUACUAACGCCAAGACCUUUCCUGGAUGA ------------..(((..(((.(((((.......))))))))..)))(((((............)))))....((......)).... ( -22.70, z-score = -2.07, R) >droEre2.scaffold_4784 371781 75 - 25762168 -------------UGUGGCGCCCAGACUAAAGCCCAGUCUGGCAACACGGCGUAAACAAGUACUAACGCCAAGACCUUUCCUGGAGGA -------------.(((..(((.(((((.......))))))))..)))(((((............)))))....((((.....)))). ( -24.50, z-score = -1.64, R) >consensus _____________UGUGGCGCCCAGACUAAAGCCCAGUCUGGCAACACGGCGAAAACAAGUACUAACGCCAAGACCUUUCCUGGAGGA ..............(((..(((.(((((.......))))))))..)))((((..............))))....((((.....)))). (-22.49 = -22.49 + 0.00)
| Location | 891,155 – 891,225 |
|---|---|
| Length | 70 |
| Sequences | 6 |
| Columns | 72 |
| Reading direction | forward |
| Mean pairwise identity | 74.41 |
| Shannon entropy | 0.49578 |
| G+C content | 0.51067 |
| Mean single sequence MFE | -17.45 |
| Consensus MFE | -7.43 |
| Energy contribution | -8.48 |
| Covariance contribution | 1.06 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.74 |
| Structure conservation index | 0.43 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.28 |
| SVM RNA-class probability | 0.624102 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
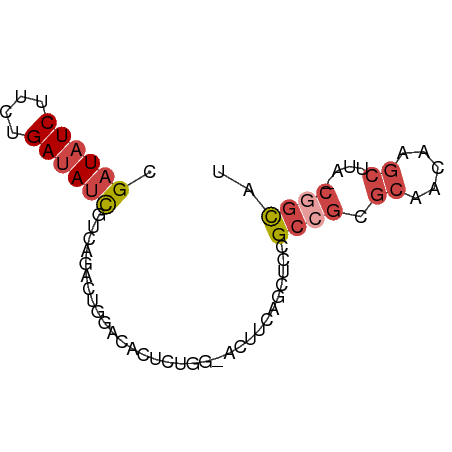
>dm3.chr2RHet 891155 70 + 3288761 CGAUAUCUUCUGAUAUCGUCAGACUGGACACACUGAAACUUCAGCUCCGC-GCGCAACAAGCUUA-CGGCAU .((((((....))))))(((.....(((....((((....)))).)))..-((.......))...-.))).. ( -16.00, z-score = -1.25, R) >droAna3.scaffold_13250 842571 67 - 3535662 CGAUAUCGUAUGAGAGUAUCGAUAUCGCUUUUUCUGCGCCUCA----CGACAU-CGACAGCCUCAUCCGCAG (((((((((((....))).))))))))......(((((.....----.(....-)((.....))...))))) ( -16.00, z-score = -0.85, R) >droSim1.chrU 4942561 69 - 15797150 CGAUAUCUUCUGAUAUCGUCAGACUGGACACUACAA--CUUCAACUCCGCCGCGCAACAAGCUUA-CGGUAU .((((((....))))))(((......))).......--..........((((.((.....))...-)))).. ( -14.60, z-score = -1.88, R) >droYak2.chrU 2414669 70 + 28119190 CGAUAUCUUCUGAUAUCGUCAGACUGGCCACUCUGG-ACUUCAACCCCGCCGCGCAACAAGCUUA-CGGCAU (((((((....)))))))(((((........)))))-...........((((.((.....))...-)))).. ( -18.60, z-score = -1.99, R) >droEre2.scaffold_4784 371856 70 - 25762168 CGAUAUCUUUUGAUAUCUUCAGACUGGCCACUCUGG-ACUCCAGCCUUGCCGCGCAACAAGCUUA-CGGCAU .((((((....))))))...((.((((((.....))-...)))).))(((((.((.....))...-))))). ( -19.70, z-score = -2.35, R) >droSec1.super_94 45144 71 - 93855 CGAUAUCUUCUGAUAUCGUCAGACUGGACACACUGGUACUUCAGCUCUGCCGCGCAACAAGCUUA-CGGCAC (((((((....)))))))..((((((((.((....))..))))).)))((((.((.....))...-)))).. ( -19.80, z-score = -2.14, R) >consensus CGAUAUCUUCUGAUAUCGUCAGACUGGACACUCUGG_ACUUCAGCUCCGCCGCGCAACAAGCUUA_CGGCAU .((((((....))))))...............................((((.((.....))....)))).. ( -7.43 = -8.48 + 1.06)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:59:24 2011