
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 22,783,526 – 22,783,628 |
| Length | 102 |
| Max. P | 0.884779 |

| Location | 22,783,526 – 22,783,628 |
|---|---|
| Length | 102 |
| Sequences | 9 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 54.85 |
| Shannon entropy | 0.97209 |
| G+C content | 0.38876 |
| Mean single sequence MFE | -18.09 |
| Consensus MFE | -7.30 |
| Energy contribution | -7.57 |
| Covariance contribution | 0.26 |
| Combinations/Pair | 1.83 |
| Mean z-score | -0.74 |
| Structure conservation index | 0.40 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.07 |
| SVM RNA-class probability | 0.884779 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
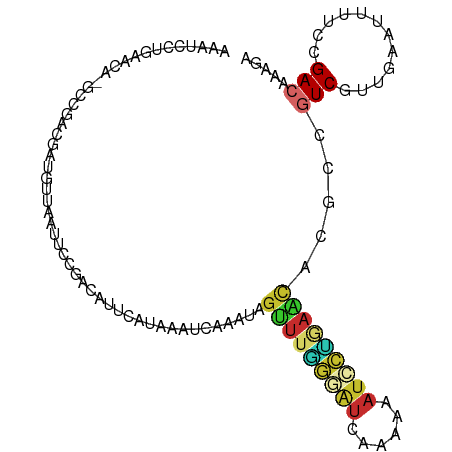
>dm3.chr2L 22783526 102 + 23011544 AAAUCCUGAACAAGCCGACGAUGUUAAUUCCGUCAUUCUUGAAUCAAAAAGUUUAGGAUCCACAAAUCCUGAGCACGCCGUCAUUGAGUUGUCCGACAAAGA ................((((((.(((((..(((((....)))........(((((((((......)))))))))....))..)))))))))))......... ( -20.20, z-score = -0.73, R) >droPer1.super_104 66497 102 + 175310 AUAUCCUGAGCACGCCGUCGUUGAAUUAUCCGACACUAAAAGUUCGAAUAGUUUGGGAUUAAAGAAUCCUGAACACGCCGUCGUUGAAUUAUCCGACAAUGA ................((((...((((...((((((.....)).......(((..(((((....)))))..))).....))))...))))...))))..... ( -22.50, z-score = -0.69, R) >dp4.Unknown_singleton_2191 8287 102 + 19051 AUAUCCUGAACCCGCCGUCGUUGAAUUAUCCGACACUAAAAGUUCGAAUAGUUUGGGAUUAAAGAAUCCUGAACACGCCGUCGUUGAAUUAUCCGACAAUGA ................((((...((((...((((((.....)).......(((..(((((....)))))..))).....))))...))))...))))..... ( -22.50, z-score = -1.11, R) >droEre2.scaffold_4845 1060992 92 + 22589142 AAAUCCCGAGAA-----GCGAUGUUCCUUCCGCUAAUCAAAAGUACACUAGUUUCGGAUUACAAAAUCCGGAACACGCCGUCGAUGAUAAUUCCGAC----- .......((..(-----(((..........))))..))............((((((((((....)))))))))).....((((..((....))))))----- ( -21.00, z-score = -1.65, R) >droYak2.chr2h_random 2690655 94 - 3774259 AAAUCCUCAACACGCCGACGAUGUUAAUUCCGUCA-------AAUAAAUCGUUUGGGAUCAAAAAAUCCUAAACACGCCGUCGUAAAAUUUUCCGACAAAG- .............((.((((..........)))).-------........(((((((((......)))))))))..)).((((..........))))....- ( -19.20, z-score = -2.40, R) >droSec1.super_88 117380 102 - 123530 AAAUCCUGAACAAGCCGACGAUGUUAAUUCCGUCAGUCUUAAAUCAAAUAGCUUGGAAUCCACAAAUACCGAGCACGCAGUCGUUGAGUUUUCCGACAAAGA .............((.(((((((.......)))).)))............((((((.((......)).))))))..)).((((..(....)..))))..... ( -18.60, z-score = -0.39, R) >droSim1.chrU 13298831 102 - 15797150 AAAUCCUGAACAAGACGAAGAUGUUAAUUCCGUCAGUCUUAAAUCAAAUAGCUUGGAAUCCACAAAUACCGAGCACGCAGUCGUUGAGUUUUCCGACAAAGA .....(((...(((((...((((.......)))).)))))..........((((((.((......)).))))))...)))..((((.......))))..... ( -19.30, z-score = -0.71, R) >anoGam1.chrU 58068247 81 + 59568033 ---------------------GAAACAAAUCAAUAUCGAUCAUUUGGAUAGUUCAAUGGCAACGGAUAUGAAAAAAAUUCUCAUUGAGUAUAAGGAUAUAUU ---------------------.......(((..(((...(((...((((..((((.((....))....))))....))))....)))..)))..)))..... ( -8.40, z-score = 0.82, R) >apiMel3.GroupUn 322208982 87 - 399230636 ---------------UUUUAAUAAUCCCAAUGAUGUUGCUAUAUAUGGAAGCAUAAAAUAUAAUGAAAAUUGAUGUGCUUUCCAGAGAUCUCCUGAUAAUUC ---------------........(((.....(((.((........((((((((((.....((((....)))).))))).))))))).)))....)))..... ( -11.10, z-score = 0.17, R) >consensus AAAUCCUGAACA_GCCGACGAUGUUAAUUCCGACAUUCAUAAAUCAAAUAGUUUGGGAUCAAAAAAUCCUGAACACGCCGUCGUUGAAUUUUCCGACAAAGA ..................................................(((((((((......))))))))).....(((............)))..... ( -7.30 = -7.57 + 0.26)


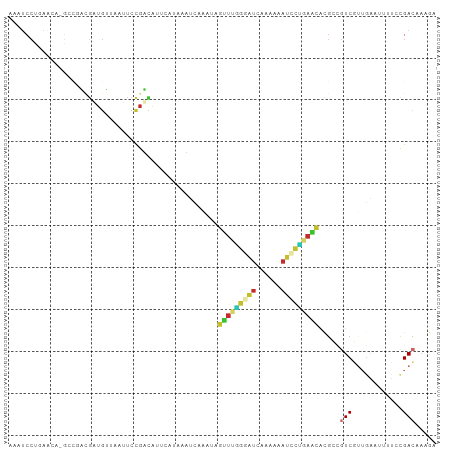
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:58:38 2011