
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 22,393,015 – 22,393,074 |
| Length | 59 |
| Max. P | 0.997681 |

| Location | 22,393,015 – 22,393,074 |
|---|---|
| Length | 59 |
| Sequences | 5 |
| Columns | 59 |
| Reading direction | forward |
| Mean pairwise identity | 84.24 |
| Shannon entropy | 0.28441 |
| G+C content | 0.64317 |
| Mean single sequence MFE | -22.68 |
| Consensus MFE | -15.58 |
| Energy contribution | -17.62 |
| Covariance contribution | 2.04 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.77 |
| Structure conservation index | 0.69 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.33 |
| SVM RNA-class probability | 0.648243 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
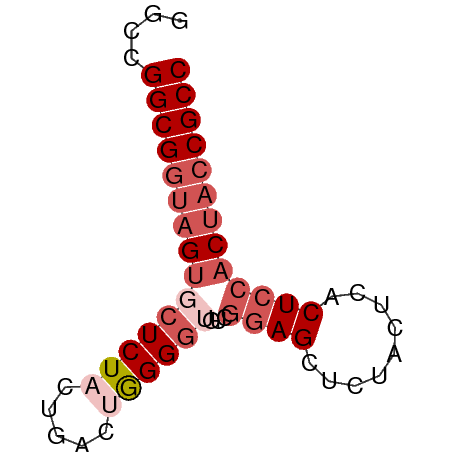
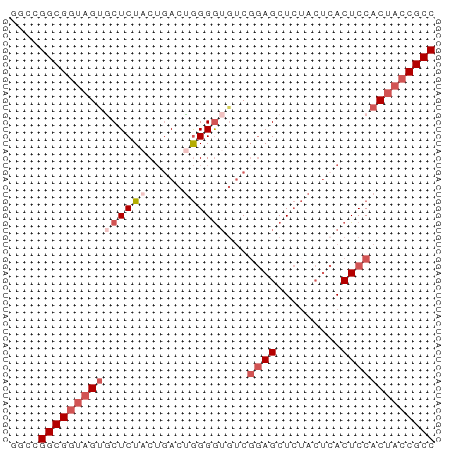
>dm3.chr2L 22393015 59 + 23011544 GGCCGGCGGUAGUGCUCUACUGACUGGGGUCUCGGAGCUCUACUCACUCCACCACCGCC ....((((((.(((......(((.((((((......)))))).)))...))).)))))) ( -20.30, z-score = -0.34, R) >droEre2.scaffold_4929 12726921 59 - 26641161 GGCCGGCGGUAGCACUCUCCUAACUGGGGAGACCGAGCUCUACUCACUCCACUACCGCC ....((((((((..((((((.....))))))...(((.....)))......)))))))) ( -26.00, z-score = -3.60, R) >droYak2.chr2L 7924854 53 + 22324452 GGCCGGCGUCAGUUCUCUCC-GACAAGGAUGUCGAAGCU-----GACUCCACUACCGCC ....((.(((((((.....(-((((....))))).))))-----))).))......... ( -16.30, z-score = -0.91, R) >droSec1.super_162 6877 59 + 37850 GGCCGGCGGUAGUGCUCUACUGACUGGGGUGUCGGAGCUCUACUCACUCCACUACCGCC ....((((((((((......(((.((((((......)))))).)))...)))))))))) ( -25.40, z-score = -2.00, R) >droSim1.chrU 11708030 59 + 15797150 GGCCGGCGGUAGUGCUCUACUGACUGGGGUGUCGGAGCUCUACUCACUCCACUACCGCC ....((((((((((......(((.((((((......)))))).)))...)))))))))) ( -25.40, z-score = -2.00, R) >consensus GGCCGGCGGUAGUGCUCUACUGACUGGGGUGUCGGAGCUCUACUCACUCCACUACCGCC ....(((((((((((((((.....))))))...((((.........))))))))))))) (-15.58 = -17.62 + 2.04)

| Location | 22,393,015 – 22,393,074 |
|---|---|
| Length | 59 |
| Sequences | 5 |
| Columns | 59 |
| Reading direction | reverse |
| Mean pairwise identity | 84.24 |
| Shannon entropy | 0.28441 |
| G+C content | 0.64317 |
| Mean single sequence MFE | -26.16 |
| Consensus MFE | -22.24 |
| Energy contribution | -22.24 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.29 |
| Mean z-score | -3.17 |
| Structure conservation index | 0.85 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 3.15 |
| SVM RNA-class probability | 0.997681 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
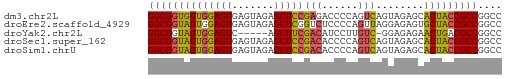
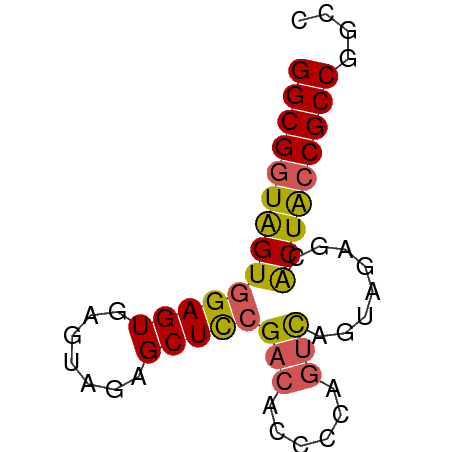
>dm3.chr2L 22393015 59 - 23011544 GGCGGUGGUGGAGUGAGUAGAGCUCCGAGACCCCAGUCAGUAGAGCACUACCGCCGGCC ((((((((((((((.......)))))..(((....)))........))))))))).... ( -26.30, z-score = -2.94, R) >droEre2.scaffold_4929 12726921 59 + 26641161 GGCGGUAGUGGAGUGAGUAGAGCUCGGUCUCCCCAGUUAGGAGAGUGCUACCGCCGGCC (((((((((.((((.......))))(.(((((.......))))).)))))))))).... ( -27.80, z-score = -3.15, R) >droYak2.chr2L 7924854 53 - 22324452 GGCGGUAGUGGAGUC-----AGCUUCGACAUCCUUGUC-GGAGAGAACUGACGCCGGCC ...(((...((.(((-----((((((((((....))))-))))....))))).)).))) ( -21.90, z-score = -1.91, R) >droSec1.super_162 6877 59 - 37850 GGCGGUAGUGGAGUGAGUAGAGCUCCGACACCCCAGUCAGUAGAGCACUACCGCCGGCC ((((((((((((((.......)))))(((......)))........))))))))).... ( -27.40, z-score = -3.94, R) >droSim1.chrU 11708030 59 - 15797150 GGCGGUAGUGGAGUGAGUAGAGCUCCGACACCCCAGUCAGUAGAGCACUACCGCCGGCC ((((((((((((((.......)))))(((......)))........))))))))).... ( -27.40, z-score = -3.94, R) >consensus GGCGGUAGUGGAGUGAGUAGAGCUCCGACACCCCAGUCAGUAGAGCACUACCGCCGGCC ((((((((((((((.......)))))(((......)))........))))))))).... (-22.24 = -22.24 + 0.00)
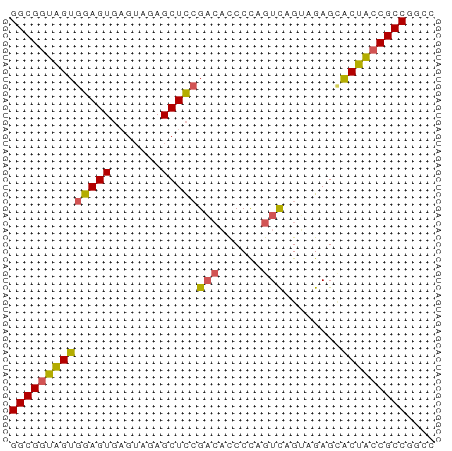
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:57:55 2011