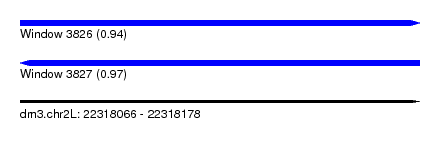

| Sequence ID | dm3.chr2L |
|---|---|
| Location | 22,318,066 – 22,318,178 |
| Length | 112 |
| Max. P | 0.968726 |

| Location | 22,318,066 – 22,318,178 |
|---|---|
| Length | 112 |
| Sequences | 3 |
| Columns | 112 |
| Reading direction | forward |
| Mean pairwise identity | 82.74 |
| Shannon entropy | 0.25630 |
| G+C content | 0.39446 |
| Mean single sequence MFE | -27.47 |
| Consensus MFE | -22.50 |
| Energy contribution | -22.50 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.99 |
| Structure conservation index | 0.82 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.40 |
| SVM RNA-class probability | 0.935832 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
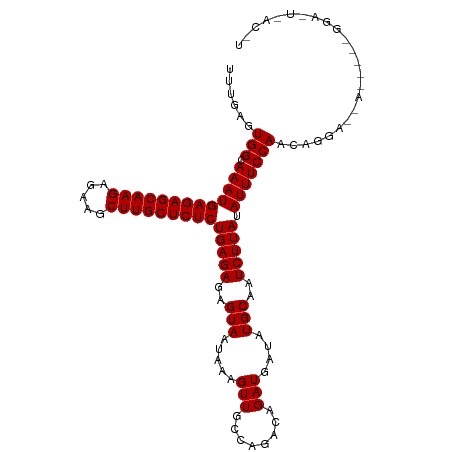
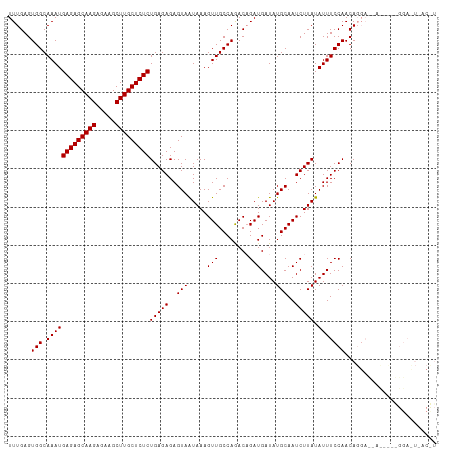
>dm3.chr2L 22318066 112 + 23011544 UUUGAGUGGCAAAUGAGAGCAAGAGAAGCUUGCUCUCUGAGAGAGUAAUAAAGUUGCCAAACAGAUGAUAUGCAAUCUUAUAUUUCCAACAGGACCACCUUGGGACUUACCU ((((..((((((..(((((((((.....)))))))))(....)..........))))))..))))..................(((((..(((....))))))))....... ( -28.90, z-score = -1.55, R) >droSim1.chr2L_random 889169 91 + 909653 UUUGAGUGGCAAAUGAGAGCAAGAGAAGCUUGCUCUCUGAGAGAGUAGUAAAGUUGCCAGACCGAUGAUAUGCAAUCUUAUAUUUCCAACA--------------------- ......(((.(((((((((((((.....)))))))))(((((..(((.....((((......))))....)))..))))).)))))))...--------------------- ( -24.60, z-score = -2.04, R) >droSec1.super_112 64358 112 - 109131 UUUGAGUGGCAAAUGAGAGCAAGAGAAGCUUGCUCUCUGAGAGAGUAAUAAAGUUGCCAGACAGAUGAUAUGCAAUCUUAUAUUUCCAACAGGAUAAUACAAGGAUUAACUU ((((..((((((..(((((((((.....)))))))))(....)..........))))))..))))........((((((.(((((((....))).))))..))))))..... ( -28.90, z-score = -2.39, R) >consensus UUUGAGUGGCAAAUGAGAGCAAGAGAAGCUUGCUCUCUGAGAGAGUAAUAAAGUUGCCAGACAGAUGAUAUGCAAUCUUAUAUUUCCAACAGGA__A_____GGA_U_AC_U ......(((.(((((((((((((.....)))))))))(((((..(((.....(((........)))....)))..))))).)))))))........................ (-22.50 = -22.50 + 0.00)

| Location | 22,318,066 – 22,318,178 |
|---|---|
| Length | 112 |
| Sequences | 3 |
| Columns | 112 |
| Reading direction | reverse |
| Mean pairwise identity | 82.74 |
| Shannon entropy | 0.25630 |
| G+C content | 0.39446 |
| Mean single sequence MFE | -26.78 |
| Consensus MFE | -21.17 |
| Energy contribution | -21.50 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.36 |
| Structure conservation index | 0.79 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.80 |
| SVM RNA-class probability | 0.968726 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
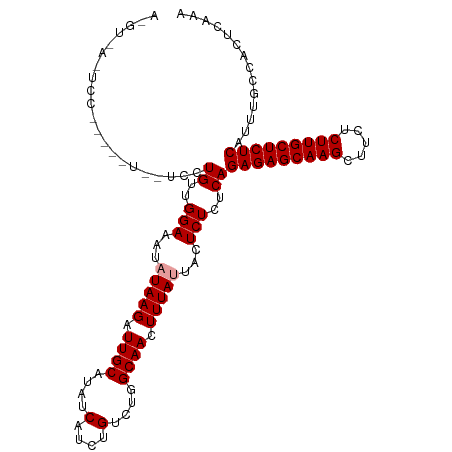
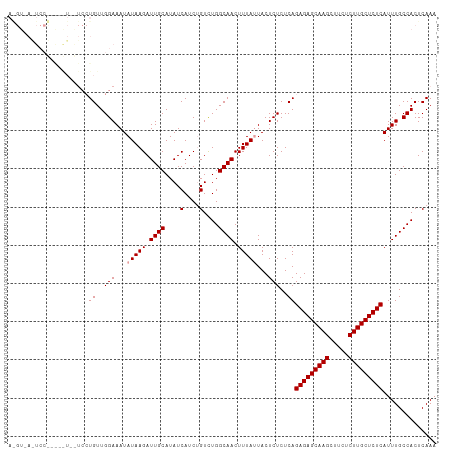
>dm3.chr2L 22318066 112 - 23011544 AGGUAAGUCCCAAGGUGGUCCUGUUGGAAAUAUAAGAUUGCAUAUCAUCUGUUUGGCAACUUUAUUACUCUCUCAGAGAGCAAGCUUCUCUUGCUCUCAUUUGCCACUCAAA .((......))..((((((..((..(((...(((((.((((....(....)....)))).)))))...)))..))(((((((((.....)))))))))....)))))).... ( -30.20, z-score = -1.83, R) >droSim1.chr2L_random 889169 91 - 909653 ---------------------UGUUGGAAAUAUAAGAUUGCAUAUCAUCGGUCUGGCAACUUUACUACUCUCUCAGAGAGCAAGCUUCUCUUGCUCUCAUUUGCCACUCAAA ---------------------..((((.......((((((........))))))(((((................(((((((((.....)))))))))..)))))..)))). ( -20.97, z-score = -1.56, R) >droSec1.super_112 64358 112 + 109131 AAGUUAAUCCUUGUAUUAUCCUGUUGGAAAUAUAAGAUUGCAUAUCAUCUGUCUGGCAACUUUAUUACUCUCUCAGAGAGCAAGCUUCUCUUGCUCUCAUUUGCCACUCAAA ..((.((((.(((((((.(((....))))))))))))))))........((..((((((................(((((((((.....)))))))))..))))))..)).. ( -29.17, z-score = -3.69, R) >consensus A_GU_A_UCC_____U__UCCUGUUGGAAAUAUAAGAUUGCAUAUCAUCUGUCUGGCAACUUUAUUACUCUCUCAGAGAGCAAGCUUCUCUUGCUCUCAUUUGCCACUCAAA .....................((..(((...(((((.((((....(....)....)))).)))))...)))..))(((((((((.....))))))))).............. (-21.17 = -21.50 + 0.33)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:57:49 2011