
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 22,273,538 – 22,273,633 |
| Length | 95 |
| Max. P | 0.961739 |

| Location | 22,273,538 – 22,273,633 |
|---|---|
| Length | 95 |
| Sequences | 10 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 49.96 |
| Shannon entropy | 0.97945 |
| G+C content | 0.36547 |
| Mean single sequence MFE | -19.00 |
| Consensus MFE | -6.49 |
| Energy contribution | -7.09 |
| Covariance contribution | 0.60 |
| Combinations/Pair | 2.00 |
| Mean z-score | -1.35 |
| Structure conservation index | 0.34 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.70 |
| SVM RNA-class probability | 0.961739 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 22273538 95 - 23011544 --UUUAAAACCGAAACGGUUAUUGCUGAAGGCAGUUU-AUAUAGUUCGGGAUCUGAA--AAUCCUGAACACGCC--GACGUUGAAUUAUCCGUCAUUAAUAA------------ --.....(((((...)))))(((((.....)))))..-.....(((((((((.....--.))))))))).....--((((..........))))........------------ ( -23.10, z-score = -1.77, R) >droSim1.chrU 10968444 95 + 15797150 --UAUAUAACCGAAACGGUAAUUGCUGAAGGCAGCUU-AUAUAGCUUGGGAUCAAUG--AAUCCUGAGCACGCC--AGCGUAGAUUUAUCCGCUAAUAAACA------------ --((((((((((...))))....((((....)))).)-)))))(((..((((.....--.))))..))).....--((((.(......).))))........------------ ( -25.00, z-score = -1.92, R) >droSec1.super_62 85869 95 - 174595 --UAUAAAACCGAAACGGUAAUUGCUGAAGGCAGUUU-AUAUAGCUUCGGAUCAAUG--AAUCCUGAGCACAAC--GACGUUGAAUUAUCCGUCGUUAAUAA------------ --......((((...))))((((((.....)))))).-.....((((.((((.....--.)))).))))..(((--((((..........))))))).....------------ ( -23.80, z-score = -2.26, R) >droYak2.chr2R 20717010 93 + 21139217 --UAUAAAACUGAAUCAGUUAUUGCAGAAAGAAGUAG-UUAUAGCUGGAGAUCUUAU--AAUCUCGAGCACGCC--GACGAUGGUAAUUCCGUCAAAUUG-------------- --.....(((((...)))))(((((........))))-)....(((.(((((.....--.))))).))).....--...(((((.....)))))......-------------- ( -20.90, z-score = -1.57, R) >droEre2.scaffold_4929 4037520 95 + 26641161 --CUUACAACAGAAUCAAUUAUUGUAGAGGAUAGUAU-AAAUAGCUCGGGAUCGAGU--UAUCCUGAGCAUACU--GGCGUAGAAAUAUCCGCCACCAAUAA------------ --.((((((............)))))).((..(((((-.....(((((((((.....--.))))))))))))))--((((((....))..)))).)).....------------ ( -24.80, z-score = -2.62, R) >dp4.Unknown_singleton_2191 10907 87 + 19051 --AUGAAAAGUGAAUCAGUAAACGUAGAAGGAAGUUU-AUAUAGCUUGGGAUCAAAA--AAUCCUGAGCACGCC--GACGAAGAAUUAUCCGUC-------------------- --.......(((.....((((((..........))))-))...(((..((((.....--.))))..))).))).--((((..........))))-------------------- ( -19.60, z-score = -2.04, R) >droPer1.super_230 12168 87 - 40990 --AUGAAAAGCGAUUCAGUAAAGGUAGAAGGAAGUUU-AUAUAGCUUGGGAUCAAAA--AAUCCUGAGCACGCC--GACGAAGAAUUAUCCGUC-------------------- --.......(((...........(((((......)))-))...(((..((((.....--.))))..))).))).--((((..........))))-------------------- ( -18.70, z-score = -1.30, R) >droAna3.scaffold_13079 85337 80 - 742402 --UAAAAAUCAGAUUCACUCAAGGUUGAAAUAAACAGCACAAAGUCCGAUGCUUCAAGUGAUAGUGUUUCUAUCAAAACUUA-------------------------------- --........(((..((((...(.((((.......(((((.......).)))))))).)...))))..)))...........-------------------------------- ( -10.91, z-score = -0.30, R) >droVir3.scaffold_13324 359556 110 - 2960039 CCUCUAAAAGAAACAUUAAAAGU--UUCCGACAAACG-GAAAACUUGAAAAUUUGAA-GAGUAGUGAAAAUACCCUGAGCUGGAAAUAUUGAUUGUUCAUAAGACUUAAAAGUG .........(((..(((((((((--(((((.....))-).))))))....((((..(-(..(((.(......).)))..))..)))).)))))..)))................ ( -13.50, z-score = 0.28, R) >triCas2.singleUn_299 7232 95 - 17920 --UUCAAAUGUAAAAUUUCAACUAGCUCCUAGAACAG-AAACAAUUGUAAAAAUUAA--AGCCAUAAACGGA----GAAAUAAAAGAAGGCAUUUGUCCAAAUA---------- --..(((((((....(((.......((((........-....((((.....))))..--..........)))----).....)))....)))))))........---------- ( -9.65, z-score = -0.05, R) >consensus __UUUAAAACCGAAACAGUAAUUGCUGAAGGAAGUUU_AUAUAGCUUGGGAUCUAAA__AAUCCUGAGCACACC__GACGUAGAAUUAUCCGUCA_UAAUAA____________ ...........................................(((((((((........)))))))))............................................. ( -6.49 = -7.09 + 0.60)


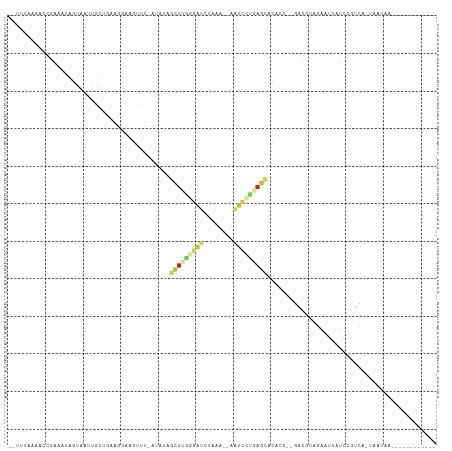
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:57:48 2011