
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 21,771,368 – 21,771,472 |
| Length | 104 |
| Max. P | 0.789505 |

| Location | 21,771,368 – 21,771,472 |
|---|---|
| Length | 104 |
| Sequences | 8 |
| Columns | 115 |
| Reading direction | forward |
| Mean pairwise identity | 73.55 |
| Shannon entropy | 0.50509 |
| G+C content | 0.46691 |
| Mean single sequence MFE | -26.57 |
| Consensus MFE | -15.95 |
| Energy contribution | -15.80 |
| Covariance contribution | -0.15 |
| Combinations/Pair | 1.50 |
| Mean z-score | -1.18 |
| Structure conservation index | 0.60 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.69 |
| SVM RNA-class probability | 0.789505 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 21771368 104 + 23011544 -------CCAUCCUUA-GGACUUUUAUCUCCAAGAGUUG--CCAGCUAAAAGGAACGACCGUAAAAGUUGAGUUAGCUGGAGUCUGAAAUUCCGGAAGCGUGUGUAGCGC-UCCG -------.........-(((.(((((.(((((.(....(--((((((....((.....)).....))))).))...))))))..))))).)))(((.((((.....))))-))). ( -29.20, z-score = -1.45, R) >droPer1.super_5 2740059 115 - 6813705 CCAUCUCUAGCUCUCACUGCCCCCAACAGAAGCCAUCUCCAAGAAGUAGAAGAAGCAACCGUAAAAGUUGAGUUAGUUCAAGUCUGGAAUUCCGGAAGCGUGUGUAGCGCAUCCA ......(((((((...(((.......)))..((..(((.(........).))).)).............)))))))......(((((....))))).((((.....))))..... ( -22.60, z-score = 0.01, R) >dp4.chr4_group1 1135132 115 + 5278887 CCAUCUCUAGCACUCACAGCCCCCAACAGAAGCCAUCUCCAAGAAGUAGAAGAAGCAACCGUAAAAGUUGAGUUAGUUCAAGUCUGGAAUUCCGGAAGCGUGUGUAGCGCAUCCA .........((.((((((((......................(((.(((......((((.......))))..))).)))...(((((....))))).)).)))).)).))..... ( -22.10, z-score = -0.29, R) >droAna3.scaffold_12943 2008389 107 - 5039921 -------UAGCUCUUAUAAGUUCCCAUCUCCUCGAGUAAAACCGGCUGAAAGAAGCGACCGUAAAAGUUGAGUUAGCUUGAGUCUGGAAUUCCGGAAGCGUGUGUAGAGC-UCCA -------.((((((((((.(((........(((((((..((((((((..................))))).))).)))))))(((((....)))))))).)))).)))))-)... ( -28.47, z-score = -1.20, R) >droEre2.scaffold_4845 19828609 100 - 22589142 -------CCAUCCUUA-GGAAUUUUAUCUCCAAGAGUUA--GCAGCUAAAAGAAGCCAACGUAAAAGUUGAGUUAGCUGCAGUCUGAAAUUCCGGAAGCGUGUAUAGCGC----- -------...(((...-(((((((((.(((...)))...--(((((((......((((((......)))).)))))))))....)))))))))))).((((.....))))----- ( -28.60, z-score = -3.02, R) >droYak2.chr2R 20065936 102 - 21139217 -------CAAUCCCUA-GGACUGUUAUCUUCAAGAGUUG--UAAGCUUAAAG--ACCAACGUAAAAGUUGAGUUAGCUGAAGUCUGAAAUUCCGGAAGUGUGUGUAGCGC-UCCG -------.........-(((.((((((((((..(((((.--((..(((...(--((((((......)))).))).....)))..)).)))))..)))).....)))))).-))). ( -24.00, z-score = -1.45, R) >droSec1.super_28 330851 104 + 873875 -------CCAUCCUUA-GGGCUUUUAUCUCCAAGAGUUG--CCAGCUAAAAGGAACGACCGUAAAAGUUGAGUUAGCUGGAGUCUGAAAUUCCGGAAGCGUGUGUAGCGC-UCCA -------.........-((..(((((.(((((.(....(--((((((....((.....)).....))))).))...))))))..)))))..))(((.((((.....))))-))). ( -28.40, z-score = -0.65, R) >droSim1.chr2L 21314276 104 + 22036055 -------CCAUCCUUA-GGACUUUUAUCUCCAAGAGUUG--CCAGCUAAAAGGAACGACCGUAAAAGUUGAGUUAGCUGGAGUCUGAAAUUCCGGAAGCGUGUGUAGCGC-UCCA -------.........-(((.(((((.(((((.(....(--((((((....((.....)).....))))).))...))))))..))))).)))(((.((((.....))))-))). ( -29.20, z-score = -1.41, R) >consensus _______CAAUCCUUA_GGACUUUUAUCUCCAAGAGUUG__CCAGCUAAAAGAAACGACCGUAAAAGUUGAGUUAGCUGGAGUCUGAAAUUCCGGAAGCGUGUGUAGCGC_UCCA .........................................((((((((....(((..........)))...)))))))).............(((.((((.....)))).))). (-15.95 = -15.80 + -0.15)


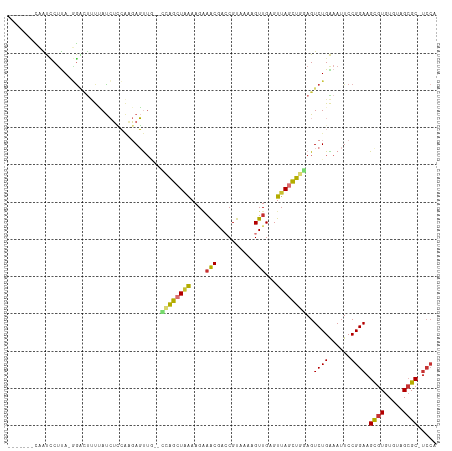
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:57:00 2011