| Sequence ID | dm3.chr2L |
|---|---|
| Location | 21,513,008 – 21,513,063 |
| Length | 55 |
| Max. P | 0.999910 |

| Location | 21,513,008 – 21,513,063 |
|---|---|
| Length | 55 |
| Sequences | 11 |
| Columns | 66 |
| Reading direction | forward |
| Mean pairwise identity | 57.00 |
| Shannon entropy | 0.92648 |
| G+C content | 0.47988 |
| Mean single sequence MFE | -19.69 |
| Consensus MFE | -12.28 |
| Energy contribution | -13.32 |
| Covariance contribution | 1.04 |
| Combinations/Pair | 1.50 |
| Mean z-score | -2.15 |
| Structure conservation index | 0.62 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 4.84 |
| SVM RNA-class probability | 0.999910 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
>dm3.chr2L 21513008 55 + 23011544 AUUGUGGCCCUGAAAAGGGCCGUUUUGGAUUAUUGUCCGCA---UUCGCAGGAGAAAA-------- .((((((((((....)))))).....((((....))))...---...)))).......-------- ( -20.10, z-score = -2.58, R) >droAna3.scaffold_13030 44769 54 - 47400 GUUGUGGCCCUGAAAAGGGCCUUUUUGGAUCAUUCUGCCCA----GCAAGCGAGCAAU-------- ((((.((((((....)))))).....(((....)))...))----))...........-------- ( -17.50, z-score = -0.99, R) >droEre2.scaffold_4845 19565920 55 - 22589142 AUUGUGGCCCUGAAGAGGGCCGUUUUGGAUUUUUGUCCGCA---CUCGCAGGAGGAAA-------- ..(((((((((....)))))).....((((....)))))))---(((....)))....-------- ( -19.20, z-score = -1.48, R) >droYak2.chr2R_random 60460 58 - 67238 AUUGUGGCCCUGAAAAGGGCCGUUUUGGAUUAAUGUCCGCAGCAUUCGUAGGAACAAA-------- .((((((((((....))))))(((.(((((....))))).)))..........)))).-------- ( -23.20, z-score = -3.62, R) >droSec1.super_91 112088 55 + 125313 AUUGUGGCCCUGAAAAGGGCCGUUUUGGAUUGUUGUCCGCA---UUCGCAGGAGCCAA-------- .....((((((....))))))(((((((((....))))((.---...)).)))))...-------- ( -21.60, z-score = -2.34, R) >droSim1.chrU 7407726 55 - 15797150 AUUGUGGCCCUGAAAAGGGCCGUUUUGGAUUGUUGUCCGCA---UUCGCAGGAGCCAA-------- .....((((((....))))))(((((((((....))))((.---...)).)))))...-------- ( -21.60, z-score = -2.34, R) >anoGam1.chr2R 47876183 55 + 62725911 UUUGUGGUCCUGAAAAGGACCGAUUUGAGAGAACGAAGCCAAUCAUUUCAGUGGC----------- (((((((((((....))))))...((....)))))))((((..........))))----------- ( -16.60, z-score = -2.43, R) >droGri2.scaffold_14978 529167 60 + 1124632 UUUGUGGCCCUGAAAAGGGCCUUUUGCUGCUAGCUGCUCGCUG------GUAUUUCUGCCUGCAAU .((((((((((....))))))......(((((((.....))))------))).........)))). ( -23.60, z-score = -2.79, R) >droMoj3.scaffold_6540 26878807 65 - 34148556 UUUGUGGCCCUGAAAAGGGCCUUUUGCUUCU-UCUUUUGGAUGUUCUGCAUACACCGAGGUAAGAU .....((((((....)))))).......(((-(.((((((.(((.......))))))))).)))). ( -22.60, z-score = -2.92, R) >droVir3.scaffold_13047 1628952 66 + 19223366 UUUGUGGCCCUGAAAAGGGCCUUUUGCUUUUGUUUGCUUCCCGUACAAUGCACAUCUAGCUGAAUU .....((((((....))))))(((.(((...((.(((............))).))..))).))).. ( -18.20, z-score = -1.84, R) >droWil1.scaffold_181036 6031 60 + 110856 AUUGUGGCCCUGAAAAGGGUUUUUUUUUUAAAAGGAAGCGUGUGUUUAAGCUAGGGACAU------ .((.(((((((....)))((((((((.....))))))))..........)))).))....------ ( -12.40, z-score = -0.37, R) >consensus AUUGUGGCCCUGAAAAGGGCCGUUUUGGAUUAUUGUCCGCA___UUCGCAGGAGCCAA________ .....((((((....))))))............................................. (-12.28 = -13.32 + 1.04)
| Location | 21,513,008 – 21,513,063 |
|---|---|
| Length | 55 |
| Sequences | 11 |
| Columns | 66 |
| Reading direction | reverse |
| Mean pairwise identity | 57.00 |
| Shannon entropy | 0.92648 |
| G+C content | 0.47988 |
| Mean single sequence MFE | -15.57 |
| Consensus MFE | -10.82 |
| Energy contribution | -10.84 |
| Covariance contribution | 0.02 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.73 |
| Structure conservation index | 0.69 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 4.35 |
| SVM RNA-class probability | 0.999768 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
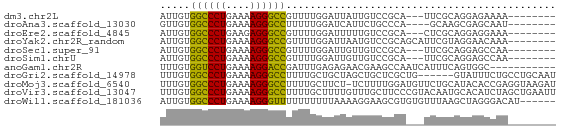
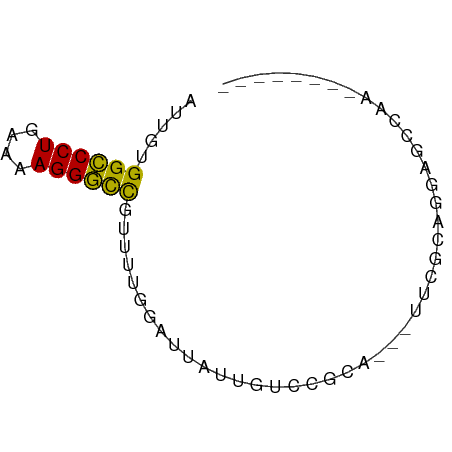
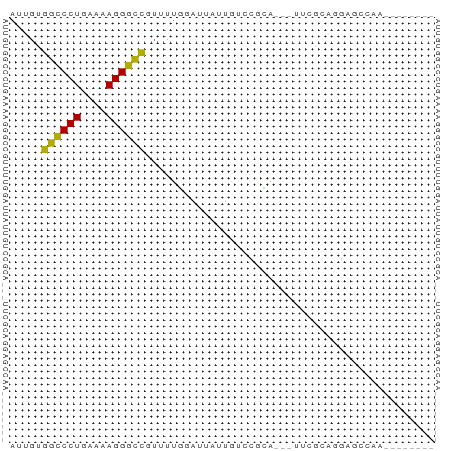
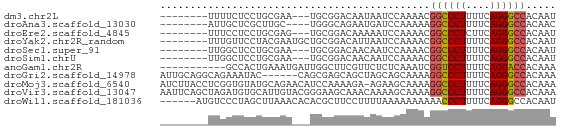
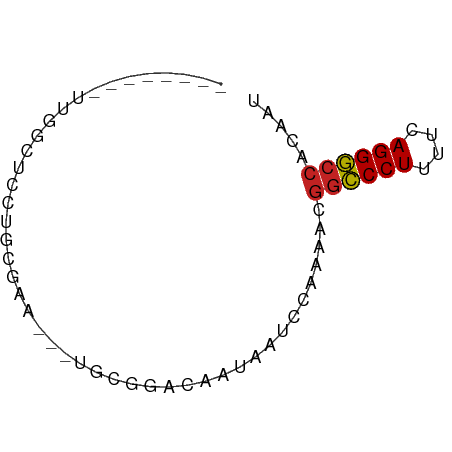
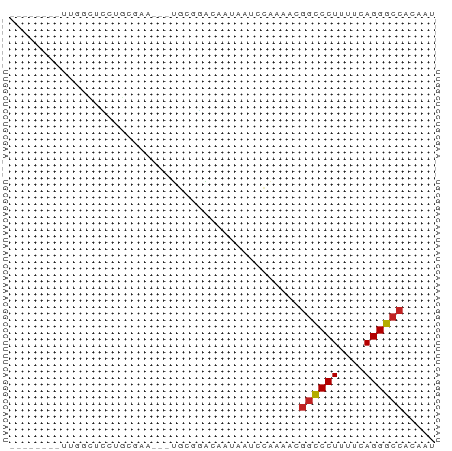
>dm3.chr2L 21513008 55 - 23011544 --------UUUUCUCCUGCGAA---UGCGGACAAUAAUCCAAAACGGCCCUUUUCAGGGCCACAAU --------.......((((...---.))))...............((((((....))))))..... ( -15.70, z-score = -2.52, R) >droAna3.scaffold_13030 44769 54 + 47400 --------AUUGCUCGCUUGC----UGGGCAGAAUGAUCCAAAAAGGCCCUUUUCAGGGCCACAAC --------.(((((((.....----))))))).............((((((....))))))..... ( -18.10, z-score = -1.59, R) >droEre2.scaffold_4845 19565920 55 + 22589142 --------UUUCCUCCUGCGAG---UGCGGACAAAAAUCCAAAACGGCCCUCUUCAGGGCCACAAU --------.......((((...---.))))...............((((((....))))))..... ( -15.70, z-score = -1.91, R) >droYak2.chr2R_random 60460 58 + 67238 --------UUUGUUCCUACGAAUGCUGCGGACAUUAAUCCAAAACGGCCCUUUUCAGGGCCACAAU --------(((((....)))))......(((......))).....((((((....))))))..... ( -15.40, z-score = -2.01, R) >droSec1.super_91 112088 55 - 125313 --------UUGGCUCCUGCGAA---UGCGGACAACAAUCCAAAACGGCCCUUUUCAGGGCCACAAU --------(((..(((.((...---.)))))...)))........((((((....))))))..... ( -16.30, z-score = -1.62, R) >droSim1.chrU 7407726 55 + 15797150 --------UUGGCUCCUGCGAA---UGCGGACAACAAUCCAAAACGGCCCUUUUCAGGGCCACAAU --------(((..(((.((...---.)))))...)))........((((((....))))))..... ( -16.30, z-score = -1.62, R) >anoGam1.chr2R 47876183 55 - 62725911 -----------GCCACUGAAAUGAUUGGCUUCGUUCUCUCAAAUCGGUCCUUUUCAGGACCACAAA -----------((((..........))))................((((((....))))))..... ( -13.50, z-score = -2.00, R) >droGri2.scaffold_14978 529167 60 - 1124632 AUUGCAGGCAGAAAUAC------CAGCGAGCAGCUAGCAGCAAAAGGCCCUUUUCAGGGCCACAAA .((((.((.(....).)------)(((.....)))....))))..((((((....))))))..... ( -20.30, z-score = -2.35, R) >droMoj3.scaffold_6540 26878807 65 + 34148556 AUCUUACCUCGGUGUAUGCAGAACAUCCAAAAGA-AGAAGCAAAAGGCCCUUUUCAGGGCCACAAA .((((.....(((((.......)))))......)-))).......((((((....))))))..... ( -16.90, z-score = -1.95, R) >droVir3.scaffold_13047 1628952 66 - 19223366 AAUUCAGCUAGAUGUGCAUUGUACGGGAAGCAAACAAAAGCAAAAGGCCCUUUUCAGGGCCACAAA ......(((...(((((...)))))...)))..............((((((....))))))..... ( -18.30, z-score = -1.74, R) >droWil1.scaffold_181036 6031 60 - 110856 ------AUGUCCCUAGCUUAAACACACGCUUCCUUUUAAAAAAAAAACCCUUUUCAGGGCCACAAU ------.((.(((((((..........)))...((((....))))..........)))).)).... ( -4.80, z-score = 0.27, R) >consensus ________UUGGCUCCUGCGAA___UGCGGACAAUAAUCCAAAACGGCCCUUUUCAGGGCCACAAU .............................................((((((....))))))..... (-10.82 = -10.84 + 0.02)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:56:35 2011