| Sequence ID | dm3.chr2L |
|---|---|
| Location | 21,502,543 – 21,502,634 |
| Length | 91 |
| Max. P | 0.997136 |

| Location | 21,502,543 – 21,502,634 |
|---|---|
| Length | 91 |
| Sequences | 9 |
| Columns | 92 |
| Reading direction | forward |
| Mean pairwise identity | 68.61 |
| Shannon entropy | 0.65115 |
| G+C content | 0.34208 |
| Mean single sequence MFE | -20.02 |
| Consensus MFE | -9.01 |
| Energy contribution | -9.52 |
| Covariance contribution | 0.51 |
| Combinations/Pair | 1.58 |
| Mean z-score | -2.38 |
| Structure conservation index | 0.45 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 3.04 |
| SVM RNA-class probability | 0.997136 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
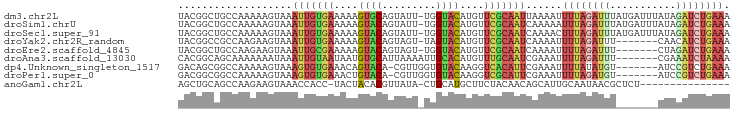
>dm3.chr2L 21502543 91 + 23011544 UACGGCUGCCAAAAAGUAAAUUGUGAAAAAGUGCAGUAUU-UGGUACAUGUUCGCAAUUAAAAUUUUAGAUUUAUGAUUUAUAGAUCUGAAA ....(((.......))).(((((((((...((((......-..))))...)))))))))....(((((((((((((...))))))))))))) ( -21.90, z-score = -2.99, R) >droSim1.chrU 5517903 91 - 15797150 UACGGCUGCCAAAAAGUAAAUUGUGAAAAAGUACAGUAUU-UGGUACAUGUUCGCAAUCAAAAAUUUAGAUUUAUGAUUUAUAGAUCUGAAA ....(((.......)))..((((((((...((((......-..))))...))))))))......((((((((((((...)))))))))))). ( -20.30, z-score = -2.76, R) >droSec1.super_91 111688 91 + 125313 UACGGCUGCCAAAAAGUAAAUUGUGAAAAAGUACAGUAUU-UGGUACAUGUUCGCAAUCAAAACUUUAGAUUUAUGAUUUAUAGAUCUGAAA ....(((.......)))..((((((((...((((......-..))))...))))))))......((((((((((((...)))))))))))). ( -20.10, z-score = -2.46, R) >droYak2.chr2R_random 60096 84 - 67238 UACGGCCGCCAAGAAGUAAAUUGUGAAAAAGUACAGUAGU-UAGUACAUGUUCGCAAUCAAAAUUUUAGAUUU-------CAACAUCUGAAA ...................((((((((...((((......-..))))...)))))))).....((((((((..-------....)))))))) ( -15.40, z-score = -1.76, R) >droEre2.scaffold_4845 19574397 84 - 22589142 UACGGCUGCCAAGAAGUAAAUUGCGAAAAAGUACAGUAGU-UGGUACAUGUUCGCAAUCAAAAUUUUAGAUUU-------CUAGAUCUGAAA (((..((....))..))).((((((((...((((......-..))))...)))))))).....(((((((((.-------...))))))))) ( -21.00, z-score = -2.80, R) >droAna3.scaffold_13030 44268 85 - 47400 CACGGCAGCAAAAAAAUAAAUUGUAAUAAUGUGCAUUAAAAUUGCACAUGUUUGCAAUCGAAAUUUUAGAUUU-------CGAAAUCUAAAA ....................((((((..(((((((.......)))))))..))))))((((((((...)))))-------)))......... ( -19.40, z-score = -2.52, R) >dp4.Unknown_singleton_1517 120 84 + 24252 GACAGCGGCCAAAAAGUAAAGUGUGAAACAGUACA-CGUUGGUGUACAAGGUCACAUUCGAAAUUUUAUAUGU-------AUCCGUCUGAAA ..((((((.((((((....(((((((....(((((-(....))))))....))))))).....))))...)).-------..))).)))... ( -20.70, z-score = -1.88, R) >droPer1.super_0 3286635 84 + 11822988 GACGGCGGCCAAAAAGUAAAGUGUGAAACUGUACA-CGUUGGUGUACAAGGUCGCAUUCGAAAUUUUAGAUGU-------AUCCGUCUGAAA ((..((((((.........(((.....)))(((((-(....))))))..))))))..))....(((((((((.-------...))))))))) ( -27.70, z-score = -3.51, R) >anoGam1.chr2L 35971851 75 - 48795086 AGCUGCAGCCAAGAAGUAAACCACC-UACUACACGUUAUA-CUGCAUGCUUCUACAACAGCAUUGCAAUAACGCUCU--------------- ....((......(.((((.......-)))).)..(((((.-.((((((((........)))).)))))))))))...--------------- ( -13.70, z-score = -0.71, R) >consensus UACGGCUGCCAAAAAGUAAAUUGUGAAAAAGUACAGUAUU_UGGUACAUGUUCGCAAUCAAAAUUUUAGAUUU_______AUAGAUCUGAAA ...................(((((((....((((.........))))....)))))))......((((((((...........)))))))). ( -9.01 = -9.52 + 0.51)
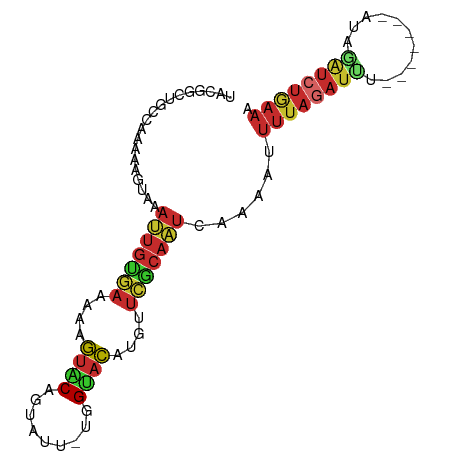
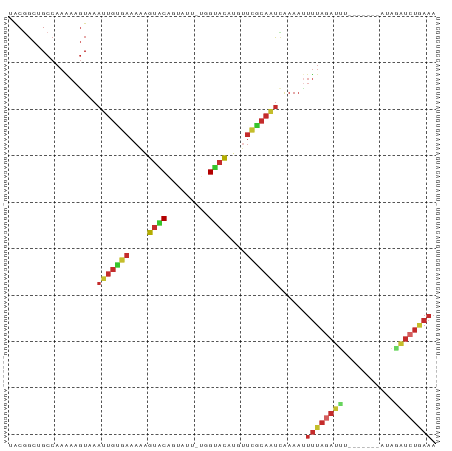
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:56:31 2011