| Sequence ID | dm3.chr2L |
|---|---|
| Location | 21,497,866 – 21,497,921 |
| Length | 55 |
| Max. P | 0.999855 |

| Location | 21,497,866 – 21,497,921 |
|---|---|
| Length | 55 |
| Sequences | 11 |
| Columns | 70 |
| Reading direction | forward |
| Mean pairwise identity | 58.18 |
| Shannon entropy | 0.84926 |
| G+C content | 0.48075 |
| Mean single sequence MFE | -19.53 |
| Consensus MFE | -12.28 |
| Energy contribution | -13.32 |
| Covariance contribution | 1.04 |
| Combinations/Pair | 1.50 |
| Mean z-score | -2.09 |
| Structure conservation index | 0.63 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 4.59 |
| SVM RNA-class probability | 0.999855 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
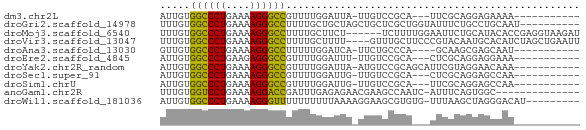
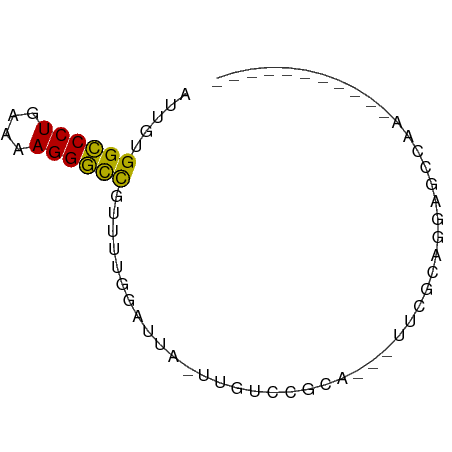
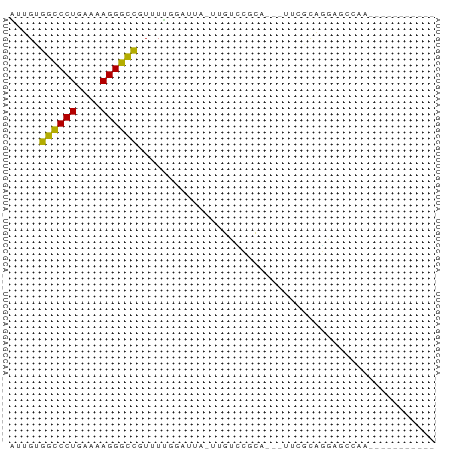
>dm3.chr2L 21497866 55 + 23011544 AUUGUGGCCCUGAAAAGGGCCGUUUUGGAUUA-UUGUCCGCA---UUCGCAGGAGAAAA----------- .((((((((((....)))))).....((((..-..))))...---...)))).......----------- ( -20.10, z-score = -2.58, R) >droGri2.scaffold_14978 515427 60 + 1124632 UUUGUGGCCCUGAAAAGGGCCUUUUGCUGCUAGCUGCUCGCUGGUAUUUCUGCCUGCAAU---------- .((((((((((....))))))......(((((((.....))))))).........)))).---------- ( -23.60, z-score = -2.79, R) >droMoj3.scaffold_6540 26857080 64 - 34148556 UUUGUGGCCCUGAAAAGGGCCUUUUGCUUCU------UCUUUUGGAAUUCUGCAUACACCGAGGUAAGAU .....((((((....))))))(((((((((.------.....((.(....).))......))))))))). ( -20.80, z-score = -2.43, R) >droVir3.scaffold_13047 1628952 66 + 19223366 UUUGUGGCCCUGAAAAGGGCCUUUUGCUUUU----GUUUGCUUCCCGUACAAUGCACAUCUAGCUGAAUU .....((((((....))))))(((.(((...----((.(((............))).))..))).))).. ( -18.20, z-score = -1.84, R) >droAna3.scaffold_13030 44769 54 - 47400 GUUGUGGCCCUGAAAAGGGCCUUUUUGGAUCA-UUCUGCCCA----GCAAGCGAGCAAU----------- ((((.((((((....)))))).....(((...-.)))...))----))...........----------- ( -17.50, z-score = -0.99, R) >droEre2.scaffold_4845 19565920 55 - 22589142 AUUGUGGCCCUGAAGAGGGCCGUUUUGGAUUU-UUGUCCGCA---CUCGCAGGAGGAAA----------- ..(((((((((....)))))).....((((..-..)))))))---(((....)))....----------- ( -19.20, z-score = -1.48, R) >droYak2.chr2R_random 60460 58 - 67238 AUUGUGGCCCUGAAAAGGGCCGUUUUGGAUUA-AUGUCCGCAGCAUUCGUAGGAACAAA----------- .((((((((((....))))))(((.(((((..-..))))).)))..........)))).----------- ( -23.20, z-score = -3.62, R) >droSec1.super_91 88087 55 + 125313 AUUGUGGCCCUGAAAAGGGCCGUUUUGGAUUG-UUGUCCGCA---CUCGCAGGAGCCAA----------- .....((((((....))))))(((((((((..-..))))((.---...)).)))))...----------- ( -21.60, z-score = -2.10, R) >droSim1.chrU 5518301 55 - 15797150 AUUGUGGCCCUGAAAAGGGCCGUUUUGGAUUG-UUGUCCGCA---UUCGCAGGAGCCAA----------- .....((((((....))))))(((((((((..-..))))((.---...)).)))))...----------- ( -21.60, z-score = -2.34, R) >anoGam1.chr2R 47876183 55 + 62725911 UUUGUGGUCCUGAAAAGGACCGAUUUGAGAGAACGAAGCCAAUC-AUUUCAGUGGC-------------- (((((((((((....))))))...((....)))))))((((...-.......))))-------------- ( -16.60, z-score = -2.43, R) >droWil1.scaffold_181036 6031 60 + 110856 AUUGUGGCCCUGAAAAGGGUUUUUUUUUUAAAAGGAAGCGUGUG-UUUAAGCUAGGGACAU--------- .((.(((((((....)))((((((((.....)))))))).....-.....)))).))....--------- ( -12.40, z-score = -0.37, R) >consensus AUUGUGGCCCUGAAAAGGGCCGUUUUGGAUUA_UUGUCCGCA___UUCGCAGGAGCCAA___________ .....((((((....))))))................................................. (-12.28 = -13.32 + 1.04)
| Location | 21,497,866 – 21,497,921 |
|---|---|
| Length | 55 |
| Sequences | 11 |
| Columns | 70 |
| Reading direction | reverse |
| Mean pairwise identity | 58.18 |
| Shannon entropy | 0.84926 |
| G+C content | 0.48075 |
| Mean single sequence MFE | -15.49 |
| Consensus MFE | -10.82 |
| Energy contribution | -10.84 |
| Covariance contribution | 0.02 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.64 |
| Structure conservation index | 0.70 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 4.06 |
| SVM RNA-class probability | 0.999595 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
>dm3.chr2L 21497866 55 - 23011544 -----------UUUUCUCCUGCGAA---UGCGGACAA-UAAUCCAAAACGGCCCUUUUCAGGGCCACAAU -----------.......((((...---.))))....-...........((((((....))))))..... ( -15.70, z-score = -2.52, R) >droGri2.scaffold_14978 515427 60 - 1124632 ----------AUUGCAGGCAGAAAUACCAGCGAGCAGCUAGCAGCAAAAGGCCCUUUUCAGGGCCACAAA ----------.((((.((.(....).))(((.....)))....))))..((((((....))))))..... ( -20.30, z-score = -2.35, R) >droMoj3.scaffold_6540 26857080 64 + 34148556 AUCUUACCUCGGUGUAUGCAGAAUUCCAAAAGA------AGAAGCAAAAGGCCCUUUUCAGGGCCACAAA ....(((....)))..(((....(((.....))------)...)))...((((((....))))))..... ( -15.10, z-score = -1.13, R) >droVir3.scaffold_13047 1628952 66 - 19223366 AAUUCAGCUAGAUGUGCAUUGUACGGGAAGCAAAC----AAAAGCAAAAGGCCCUUUUCAGGGCCACAAA ......(((...(((((...)))))...)))....----..........((((((....))))))..... ( -18.30, z-score = -1.74, R) >droAna3.scaffold_13030 44769 54 + 47400 -----------AUUGCUCGCUUGC----UGGGCAGAA-UGAUCCAAAAAGGCCCUUUUCAGGGCCACAAC -----------.(((((((.....----)))))))..-...........((((((....))))))..... ( -18.10, z-score = -1.59, R) >droEre2.scaffold_4845 19565920 55 + 22589142 -----------UUUCCUCCUGCGAG---UGCGGACAA-AAAUCCAAAACGGCCCUCUUCAGGGCCACAAU -----------.......((((...---.))))....-...........((((((....))))))..... ( -15.70, z-score = -1.91, R) >droYak2.chr2R_random 60460 58 + 67238 -----------UUUGUUCCUACGAAUGCUGCGGACAU-UAAUCCAAAACGGCCCUUUUCAGGGCCACAAU -----------(((((....)))))......(((...-...))).....((((((....))))))..... ( -15.40, z-score = -2.01, R) >droSec1.super_91 88087 55 - 125313 -----------UUGGCUCCUGCGAG---UGCGGACAA-CAAUCCAAAACGGCCCUUUUCAGGGCCACAAU -----------...((((....)))---)..(((...-...))).....((((((....))))))..... ( -17.20, z-score = -1.44, R) >droSim1.chrU 5518301 55 + 15797150 -----------UUGGCUCCUGCGAA---UGCGGACAA-CAAUCCAAAACGGCCCUUUUCAGGGCCACAAU -----------(((..(((.((...---.)))))...-)))........((((((....))))))..... ( -16.30, z-score = -1.62, R) >anoGam1.chr2R 47876183 55 - 62725911 --------------GCCACUGAAAU-GAUUGGCUUCGUUCUCUCAAAUCGGUCCUUUUCAGGACCACAAA --------------((((.......-...))))................((((((....))))))..... ( -13.50, z-score = -2.00, R) >droWil1.scaffold_181036 6031 60 - 110856 ---------AUGUCCCUAGCUUAAA-CACACGCUUCCUUUUAAAAAAAAAACCCUUUUCAGGGCCACAAU ---------.((.(((((((.....-.....)))...((((....))))..........)))).)).... ( -4.80, z-score = 0.27, R) >consensus ___________UUGGCUCCUGCGAA___UGCGGACAA_UAAUCCAAAACGGCCCUUUUCAGGGCCACAAU .................................................((((((....))))))..... (-10.82 = -10.84 + 0.02)

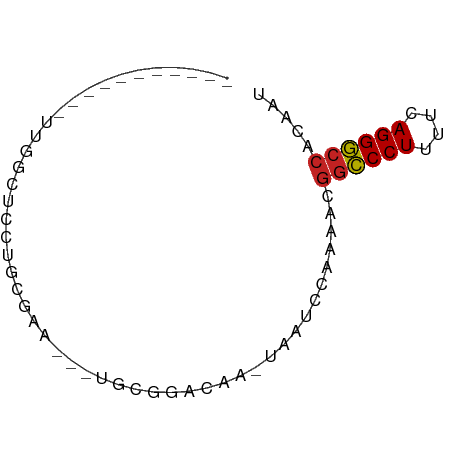
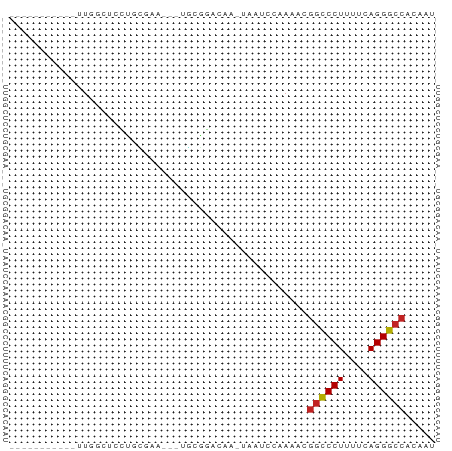
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:56:30 2011