
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 21,467,135 – 21,467,276 |
| Length | 141 |
| Max. P | 0.883039 |

| Location | 21,467,135 – 21,467,276 |
|---|---|
| Length | 141 |
| Sequences | 11 |
| Columns | 147 |
| Reading direction | forward |
| Mean pairwise identity | 62.04 |
| Shannon entropy | 0.82995 |
| G+C content | 0.29934 |
| Mean single sequence MFE | -28.90 |
| Consensus MFE | -4.82 |
| Energy contribution | -5.36 |
| Covariance contribution | 0.55 |
| Combinations/Pair | 1.33 |
| Mean z-score | -2.41 |
| Structure conservation index | 0.17 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.06 |
| SVM RNA-class probability | 0.883039 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 21467135 141 + 23011544 CAAAAAGUAAAUUGUGAA-AAAGUGCAGUAUU-UGGUACAUGUUCGCAAUUAAAAUUUUAGAUUUAUGUUUUAUAGAUCUGAAAUUUGUUUA---AACAAGUCCUUUUCAGGGCUACAACGUUCCGUU-GCAAGAGAAAAAAACUUU .........(((((((((-...((((......-..))))...))))))))).((((((((((((((((...))))))))))))))))((((.---....((((((....)))))).(((((...))))-)..........))))... ( -35.20, z-score = -3.55, R) >droSim1.chrU 1893125 141 - 15797150 CAAAAAGUAAAUUGUGAA-AAAGUACAGUAUU-UGGUACAUGUUCGCAAUCAAAAAUUUAGAUUUAUGAUUUAUAGAUCUGAAAUUUGUUUA---UACAAGUCCUUUUCAGGGCUACAACGUUCUUUU-GCAAGAGAAAAAAACUUU ......(((.((((((((-...((((......-..))))...))))))))((((..((((((((((((...)))))))))))).))))....---))).((((((....))))))......((((((.-...))))))......... ( -30.10, z-score = -2.36, R) >droSec1.super_91 82787 141 + 125313 CAAAAAGUAAAUUGUGAA-AAAGUACAGUAUU-UGGUACAUGUUCGCAAUCAAAACUUUAGAUUUAUGAUUUAUAGAUCUGAAAUUUGUUUA---UACAAGUCCUUUUCAGGGCUACAACGUUCUGUU-GCAAGAGAAAAAAACUUU ...(((((..((((((((-...((((......-..))))...))))))))......((((((((((((...))))))))))))((((((...---.)))))).(((((((((((......))))))..-..)))))......))))) ( -29.60, z-score = -1.83, R) >droYak2.chr2R_random 60105 133 - 67238 CAAGAAGUAAAUUGUGAA-AAAGUACAGUAGU-UAGUACAUGUUCGCAAUCAAAAUUUUAGAUU-------UCAACAUCUGAAAUUUGUUUA---AACAAGCCCUUUUCAGGGCUACAACGAUUCGUU-GCAAGAGAAAAAACUUU- ...(((((...(((((((-...((((......-..))))...)))))))((.(((((((((((.-------.....))))))))))).....---....((((((....)))))).(((((...))))-)...))......)))))- ( -32.50, z-score = -3.43, R) >droEre2.scaffold_4845 19565555 123 - 22589142 CAAGAAGUAAAUUGCGAA-AAAGUACAGUAGU-UGGUACAUGUUCGCAAUCAAAA----------------UAUAUAUCUGAAAUUUGUUUA---AACAAGCCCUUUUCAGGGCUACAACAUUCACUU-GCAAGAGAAAAAAAAU-- ((((......((((((((-...((((......-..))))...)))))))).....----------------........((((..((((...---....((((((....))))))))))..)))))))-)...............-- ( -26.21, z-score = -2.56, R) >droAna3.scaffold_12943 4888478 135 - 5039921 GAAAAAAUAAAUUGUGAU-AAUGUGCAUUAAAAUUGCACACGUUUGCAAUCGAAAUUUUAGAUU-------UCGAAAUCUAAAAUUAAAUUA---AACAAGCCCUUUUCAGGACUACAAGA-UAUGUUUUAAAGAGAAAGAAAUUAU ..........((((..(.-..((((((.......))))))...)..))))...(((((((((((-------....)))))))))))......---........(((((.(((((.......-...))))).)))))........... ( -24.70, z-score = -2.39, R) >dp4.Unknown_singleton_1517 15533 135 + 24252 CAAAAAGUAAAGUGUGAA-ACAGUACACGUUG-GUGUACAAGGUCGCAUUCGAAAUUUUAGAUG-------UAUCCGUCUGAAAUUGAUUUA---AACAAGCCCUUUUCAGGGCUACAAAACUAUAUUCAAAAGAGAUACAAAUUAU ......(((.(((((((.-...((((((....-))))))....))))))).(((((((((((((-------....)))))))))))......---....((((((....))))))............))........)))....... ( -34.60, z-score = -3.74, R) >droPer1.super_0 3274818 135 + 11822988 CAAAAAGUAAAGUGUGAA-ACUGUACACGUUG-GUGUACAAGGUCGCAUUCGAAAUUUUAGAUG-------UAUCCGUCUGAAAUUGAUUUA---AACAAGCCCUUUUCAGGGCUACAAAACUAUAUUCAAAAGAGAUACAAAUUAU ......(((.(((((((.-..(((((((....-)))))))...))))))).(((((((((((((-------....)))))))))))......---....((((((....))))))............))........)))....... ( -36.30, z-score = -4.06, R) >droWil1.scaffold_181036 31085 135 + 110856 CAAGAAGUGAAUAAUUAAUACUUUAUUGCAAU----CGUAAUAUUGAA---AAAAUUUUAGAUAUCGAAUACGAAAAUCUAAAAUUUUACUU--AAACACAGCCAGGUCGGCGCGUUAAGUUUCAGCG-UUAAGCGUUAAGUUUU-- .((((..(((....(((((....((((((...----.))))))(((((---((((((((((((.(((....)))..))))))))))))((((--((.....(((.....)))...))))))))))).)-))))...)))..))))-- ( -26.60, z-score = -1.36, R) >droVir3.scaffold_13047 1628447 122 + 19223366 ----GAAGAAAUAAUUUA-AAAUUGC--UAAC----UGCUUGUCAGCAUAUGCAAAUU-------------UCUAAUAAUGAAAUUUGUUUUCCUACAAAGCCCUUUUAAGGGCUACAAAUACGAUUUUCUAAGAGAAAGAAUUUU- ----...((((.......-....(((--(.((----.....)).))))...(((((((-------------((.......)))))))))))))......((((((....))))))..........(((((.....)))))......- ( -24.20, z-score = -1.87, R) >apiMel3.Group1 12264768 133 - 25854376 -AAUAUAUAAAAUAUUAG-AAAUUAAAGUAUU---UAGCGAAUUUACAUCAUUGAUCGUGACAUCGAAUGUUCCUCAUAUGAACUGCGCAAG---UCAAAGCACUGCAAAACACCGCGGAAAACAAUU----AGAAAAAUGAUUU-- -........(((((((..-.......))))))---)...........((((((..((.((((..((...((((.......))))..))...)---))).....((((........)))).........----.))..))))))..-- ( -17.90, z-score = 0.60, R) >consensus CAAAAAGUAAAUUGUGAA_AAAGUACAGUAUU_UGGUACAUGUUCGCAAUCAAAAUUUUAGAUU_______UAUACAUCUGAAAUUUGUUUA___AACAAGCCCUUUUCAGGGCUACAACAUUCUGUU_GCAAGAGAAAAAAAUUUU ...................................................................................................((((((....))))))................................ ( -4.82 = -5.36 + 0.55)

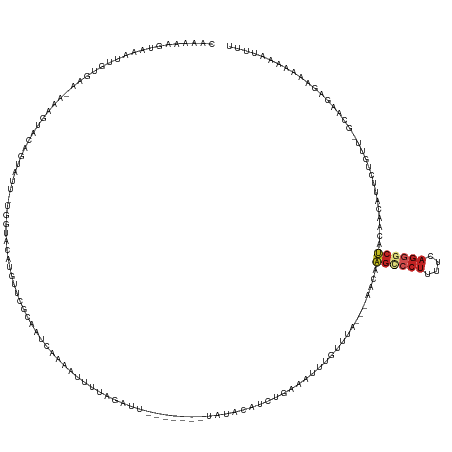

| Location | 21,467,135 – 21,467,276 |
|---|---|
| Length | 141 |
| Sequences | 11 |
| Columns | 147 |
| Reading direction | reverse |
| Mean pairwise identity | 62.04 |
| Shannon entropy | 0.82995 |
| G+C content | 0.29934 |
| Mean single sequence MFE | -27.77 |
| Consensus MFE | -2.65 |
| Energy contribution | -2.28 |
| Covariance contribution | -0.37 |
| Combinations/Pair | 2.00 |
| Mean z-score | -2.57 |
| Structure conservation index | 0.10 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.44 |
| SVM RNA-class probability | 0.695276 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 21467135 141 - 23011544 AAAGUUUUUUUCUCUUGC-AACGGAACGUUGUAGCCCUGAAAAGGACUUGUU---UAAACAAAUUUCAGAUCUAUAAAACAUAAAUCUAAAAUUUUAAUUGCGAACAUGUACCA-AAUACUGCACUUU-UUCACAAUUUACUUUUUG .(((((((((((..((((-((((...))))))))....)))))))))))...---.....((((((.((((.(((.....))).)))).)))))).(((((.(((..((((...-.....))))....-))).)))))......... ( -24.40, z-score = -1.88, R) >droSim1.chrU 1893125 141 + 15797150 AAAGUUUUUUUCUCUUGC-AAAAGAACGUUGUAGCCCUGAAAAGGACUUGUA---UAAACAAAUUUCAGAUCUAUAAAUCAUAAAUCUAAAUUUUUGAUUGCGAACAUGUACCA-AAUACUGUACUUU-UUCACAAUUUACUUUUUG .(((((((((((..((((-((.......))))))....)))))))))))(((---.....((((((.((((.(((.....))).))))))))))..(((((.(((...((((..-......))))...-))).))))))))...... ( -23.00, z-score = -1.34, R) >droSec1.super_91 82787 141 - 125313 AAAGUUUUUUUCUCUUGC-AACAGAACGUUGUAGCCCUGAAAAGGACUUGUA---UAAACAAAUUUCAGAUCUAUAAAUCAUAAAUCUAAAGUUUUGAUUGCGAACAUGUACCA-AAUACUGUACUUU-UUCACAAUUUACUUUUUG .(((((((((((..((((-(((.....)))))))....)))))))))))...---............((((.(((.....))).))))(((((...(((((.(((...((((..-......))))...-))).))))).)))))... ( -28.10, z-score = -2.59, R) >droYak2.chr2R_random 60105 133 + 67238 -AAAGUUUUUUCUCUUGC-AACGAAUCGUUGUAGCCCUGAAAAGGGCUUGUU---UAAACAAAUUUCAGAUGUUGA-------AAUCUAAAAUUUUGAUUGCGAACAUGUACUA-ACUACUGUACUUU-UUCACAAUUUACUUCUUG -...((((...(...(((-((((...)))))))(((((....)))))..)..---.))))((((((.((((.....-------.)))).)))))).(((((.(((...((((..-......))))...-))).)))))......... ( -31.40, z-score = -3.51, R) >droEre2.scaffold_4845 19565555 123 + 22589142 --AUUUUUUUUCUCUUGC-AAGUGAAUGUUGUAGCCCUGAAAAGGGCUUGUU---UAAACAAAUUUCAGAUAUAUA----------------UUUUGAUUGCGAACAUGUACCA-ACUACUGUACUUU-UUCGCAAUUUACUUCUUG --................-(((((((((((..((((((....))))))....---..))))....(((((......----------------.)))))(((((((...((((..-......))))...-)))))))))))))).... ( -30.40, z-score = -3.92, R) >droAna3.scaffold_12943 4888478 135 + 5039921 AUAAUUUCUUUCUCUUUAAAACAUA-UCUUGUAGUCCUGAAAAGGGCUUGUU---UAAUUUAAUUUUAGAUUUCGA-------AAUCUAAAAUUUCGAUUGCAAACGUGUGCAAUUUUAAUGCACAUU-AUCACAAUUUAUUUUUUC .................((((.(((-..((((.(((((....)))))((((.---......(((((((((((....-------)))))))))))......))))..(((((((.......))))))).-...))))..))).)))). ( -28.02, z-score = -3.79, R) >dp4.Unknown_singleton_1517 15533 135 - 24252 AUAAUUUGUAUCUCUUUUGAAUAUAGUUUUGUAGCCCUGAAAAGGGCUUGUU---UAAAUCAAUUUCAGACGGAUA-------CAUCUAAAAUUUCGAAUGCGACCUUGUACAC-CAACGUGUACUGU-UUCACACUUUACUUUUUG ......(((((((..((((((....((((...((((((....))))))....---.))))....)))))).)))))-------))..............((.((....((((((-....))))))...-.)).))............ ( -31.70, z-score = -3.24, R) >droPer1.super_0 3274818 135 - 11822988 AUAAUUUGUAUCUCUUUUGAAUAUAGUUUUGUAGCCCUGAAAAGGGCUUGUU---UAAAUCAAUUUCAGACGGAUA-------CAUCUAAAAUUUCGAAUGCGACCUUGUACAC-CAACGUGUACAGU-UUCACACUUUACUUUUUG ......(((((((..((((((....((((...((((((....))))))....---.))))....)))))).)))))-------))..............((.((..((((((((-....)))))))).-.)).))............ ( -34.50, z-score = -4.02, R) >droWil1.scaffold_181036 31085 135 - 110856 --AAAACUUAACGCUUAA-CGCUGAAACUUAACGCGCCGACCUGGCUGUGUUU--AAGUAAAAUUUUAGAUUUUCGUAUUCGAUAUCUAAAAUUUU---UUCAAUAUUACG----AUUGCAAUAAAGUAUUAAUUAUUCACUUCUUG --.....((((..(((..-.((....((((((((((((.....))).))).))--))))((((((((((((..(((....))).))))))))))))---............----...))....)))..)))).............. ( -26.40, z-score = -2.35, R) >droVir3.scaffold_13047 1628447 122 - 19223366 -AAAAUUCUUUCUCUUAGAAAAUCGUAUUUGUAGCCCUUAAAAGGGCUUUGUAGGAAAACAAAUUUCAUUAUUAGA-------------AAUUUGCAUAUGCUGACAAGCA----GUUA--GCAAUUU-UAAAUUAUUUCUUC---- -...............(((((.((.(((....((((((....))))))..))).))...((((((((.......))-------------))))))....(((((((.....----))))--)))....-.......)))))..---- ( -28.60, z-score = -2.95, R) >apiMel3.Group1 12264768 133 + 25854376 --AAAUCAUUUUUCU----AAUUGUUUUCCGCGGUGUUUUGCAGUGCUUUGA---CUUGCGCAGUUCAUAUGAGGAACAUUCGAUGUCACGAUCAAUGAUGUAAAUUCGCUA---AAUACUUUAAUUU-CUAAUAUUUUAUAUAUU- --.............----...(((((...((((...((((((((((.....---...))))...((((.((((..((((...))))..)..))))))))))))).)))).)---)))).........-.................- ( -19.00, z-score = 1.27, R) >consensus AAAAUUUUUUUCUCUUGC_AACAGAAUGUUGUAGCCCUGAAAAGGGCUUGUU___UAAACAAAUUUCAGAUGUAUA_______AAUCUAAAAUUUUGAUUGCGAACAUGUACCA_AAUACUGUACUUU_UUCACAAUUUACUUUUUG ................................((.(((....))).))................................................................................................... ( -2.65 = -2.28 + -0.37)

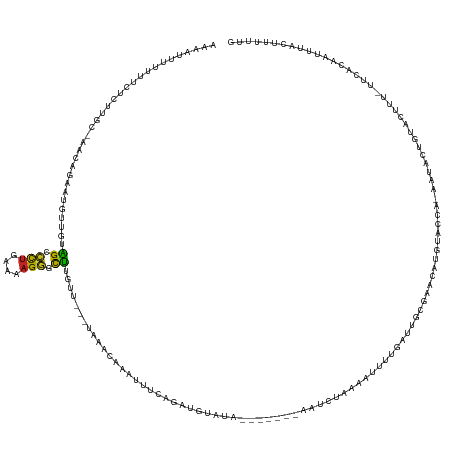

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:56:24 2011