
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 21,232,542 – 21,232,608 |
| Length | 66 |
| Max. P | 0.773842 |

| Location | 21,232,542 – 21,232,608 |
|---|---|
| Length | 66 |
| Sequences | 10 |
| Columns | 86 |
| Reading direction | forward |
| Mean pairwise identity | 69.70 |
| Shannon entropy | 0.57293 |
| G+C content | 0.39678 |
| Mean single sequence MFE | -15.70 |
| Consensus MFE | -3.71 |
| Energy contribution | -3.81 |
| Covariance contribution | 0.10 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.67 |
| Structure conservation index | 0.24 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.65 |
| SVM RNA-class probability | 0.773842 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
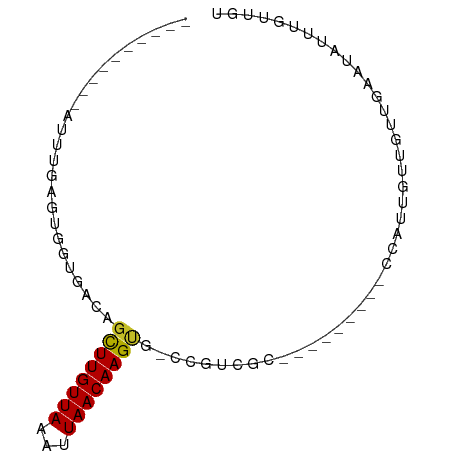
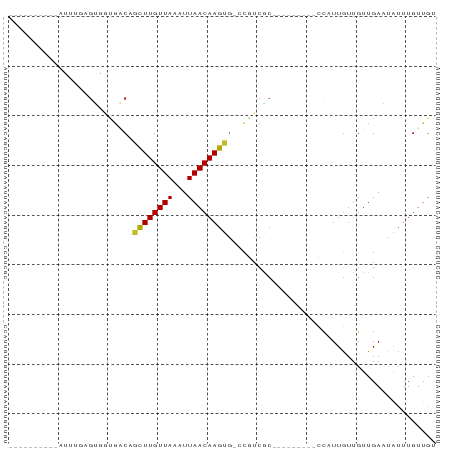
>dm3.chr2L 21232542 66 + 23011544 ACAACAAAUAUUCAACAACAAUGG---------GCGACGG-CACUUGUUAAUUUAACAAGCUGUCACCACUCAAAU---------- .......................(---------(.(((((-(..((((((...)))))))))))).))........---------- ( -14.80, z-score = -3.86, R) >droSim1.chr2L 20817135 66 + 22036055 ACAACAAAUAUUCAACAACAAUGG---------GUGACGG-UACUUGUUAAUUUAACAAGCUGUCACCACUCAAAG---------- .......................(---------(((((((-..(((((((...)))))))))))))))........---------- ( -17.90, z-score = -4.24, R) >droSec1.super_18 1157347 66 + 1342155 ACAACAAAUAUUCAACAACAAUGG---------GCGACGG-UACUUGUUAAUUUAACAAGCUGUCACCACUCCAAU---------- .......................(---------(.(((((-..(((((((...)))))))))))).))........---------- ( -14.00, z-score = -2.93, R) >droYak2.chr2R 19596855 65 - 21139217 ACAACAAAUUUUCAACAACAAUGG---------GCGGCG--CACUUGUUAAUUUAACAAGCUGUCACCACGCAAGU---------- .......................(---------(.((((--..(((((((...))))))).)))).))((....))---------- ( -12.70, z-score = -1.16, R) >droEre2.scaffold_4845 19323386 66 - 22589142 ACAACAAAUAUUCAACAACAAUGG---------GCGACGG-CACUUGUUAAUUUAACAAGCUGUCACCACGCAAAU---------- .......................(---------(.(((((-(..((((((...)))))))))))).))........---------- ( -14.80, z-score = -3.20, R) >droAna3.scaffold_12916 10971511 56 + 16180835 ACAACAA-CAUUCAACAAUAAUGA---------GUGGCAGUCACUUGUUAAUUUAACAAGCUGCCG-------------------- .......-.(((((.......)))---------))(((((...(((((((...)))))))))))).-------------------- ( -14.10, z-score = -3.12, R) >dp4.chr4_group4 3319624 84 + 6586962 ACAACAAAUAAUCAACAACAAUGAAGA--GGGAGAGAAGGCCGCUUGUUAAUUUAACAAGCGGGCGAGUCGAUCCUCCCCUCCCGU .........................((--(((.(((..(.((((((((((...)))))))))).)((.....)))))))))).... ( -28.30, z-score = -3.94, R) >droWil1.scaffold_180772 7226497 73 + 8906247 ACAACAAAAUAUCAACAACAAAGAUGAAGGGGGGAAAUGAUCGCUUGUUAAUUUAACAAGUCGCAUAUCCUGU------------- .........((((.........)))).....((((.(((...((((((((...))))))))..))).))))..------------- ( -14.00, z-score = -1.76, R) >droMoj3.scaffold_6500 18521002 67 - 32352404 ACAACAGAUGAUCAACAACAAGGU--------GCCGGAGGUCACUUGUUAAUUUAACAAGC-GGUGCUCCAAAGAU---------- .....................((.--------.((((....).(((((((...))))))))-))..))........---------- ( -11.90, z-score = 0.04, R) >droGri2.scaffold_15252 14176818 68 - 17193109 ACAACAAAUGAUCAACAACAAAAUA-------GGCGGAGGCCACUUGUUAAUUUAACAAAC-GGCUCUCCUGAGAU---------- .........................-------...(((((((..((((((...))))))..-))).))))......---------- ( -14.50, z-score = -2.48, R) >consensus ACAACAAAUAUUCAACAACAAUGG_________GCGACGG_CACUUGUUAAUUUAACAAGCUGUCACCACUCAAAU__________ ...........................................(((((((...))))))).......................... ( -3.71 = -3.81 + 0.10)



| Location | 21,232,542 – 21,232,608 |
|---|---|
| Length | 66 |
| Sequences | 10 |
| Columns | 86 |
| Reading direction | reverse |
| Mean pairwise identity | 69.70 |
| Shannon entropy | 0.57293 |
| G+C content | 0.39678 |
| Mean single sequence MFE | -17.20 |
| Consensus MFE | -5.67 |
| Energy contribution | -5.60 |
| Covariance contribution | -0.07 |
| Combinations/Pair | 1.25 |
| Mean z-score | -1.96 |
| Structure conservation index | 0.33 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.35 |
| SVM RNA-class probability | 0.656440 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 21232542 66 - 23011544 ----------AUUUGAGUGGUGACAGCUUGUUAAAUUAACAAGUG-CCGUCGC---------CCAUUGUUGUUGAAUAUUUGUUGU ----------......(.((((((.((((((((...)))))))).-..)))))---------))...................... ( -16.70, z-score = -2.42, R) >droSim1.chr2L 20817135 66 - 22036055 ----------CUUUGAGUGGUGACAGCUUGUUAAAUUAACAAGUA-CCGUCAC---------CCAUUGUUGUUGAAUAUUUGUUGU ----------......(.((((((.((((((((...)))))))).-..)))))---------))...................... ( -16.00, z-score = -2.15, R) >droSec1.super_18 1157347 66 - 1342155 ----------AUUGGAGUGGUGACAGCUUGUUAAAUUAACAAGUA-CCGUCGC---------CCAUUGUUGUUGAAUAUUUGUUGU ----------......(.((((((.((((((((...)))))))).-..)))))---------))...................... ( -15.60, z-score = -1.61, R) >droYak2.chr2R 19596855 65 + 21139217 ----------ACUUGCGUGGUGACAGCUUGUUAAAUUAACAAGUG--CGCCGC---------CCAUUGUUGUUGAAAAUUUGUUGU ----------...((.((((((...((((((((...)))))))).--))))))---------.))..................... ( -17.60, z-score = -1.65, R) >droEre2.scaffold_4845 19323386 66 + 22589142 ----------AUUUGCGUGGUGACAGCUUGUUAAAUUAACAAGUG-CCGUCGC---------CCAUUGUUGUUGAAUAUUUGUUGU ----------......(.((((((.((((((((...)))))))).-..)))))---------))...................... ( -16.80, z-score = -1.91, R) >droAna3.scaffold_12916 10971511 56 - 16180835 --------------------CGGCAGCUUGUUAAAUUAACAAGUGACUGCCAC---------UCAUUAUUGUUGAAUG-UUGUUGU --------------------.((((((((((((...)))))))...)))))..---------(((.......)))...-....... ( -13.10, z-score = -1.31, R) >dp4.chr4_group4 3319624 84 - 6586962 ACGGGAGGGGAGGAUCGACUCGCCCGCUUGUUAAAUUAACAAGCGGCCUUCUCUCCC--UCUUCAUUGUUGUUGAUUAUUUGUUGU ..(((((((((((..........((((((((((...)))))))))))))))))))))--........................... ( -31.90, z-score = -3.72, R) >droWil1.scaffold_180772 7226497 73 - 8906247 -------------ACAGGAUAUGCGACUUGUUAAAUUAACAAGCGAUCAUUUCCCCCCUUCAUCUUUGUUGUUGAUAUUUUGUUGU -------------((((((((..((.(((((((...)))))))))..............(((.(......).)))))))))))... ( -11.50, z-score = -0.94, R) >droMoj3.scaffold_6500 18521002 67 + 32352404 ----------AUCUUUGGAGCACC-GCUUGUUAAAUUAACAAGUGACCUCCGGC--------ACCUUGUUGUUGAUCAUCUGUUGU ----------......((((...(-((((((((...)))))))))..))))(((--------(...((........))..)))).. ( -15.00, z-score = -1.16, R) >droGri2.scaffold_15252 14176818 68 + 17193109 ----------AUCUCAGGAGAGCC-GUUUGUUAAAUUAACAAGUGGCCUCCGCC-------UAUUUUGUUGUUGAUCAUUUGUUGU ----------......((((.(((-((((((((...)))))))))))))))...-------......................... ( -17.80, z-score = -2.74, R) >consensus __________AUUUGAGUGGUGACAGCUUGUUAAAUUAACAAGUG_CCGUCGC_________CCAUUGUUGUUGAAUAUUUGUUGU .........................((((((((...)))))))).......................................... ( -5.67 = -5.60 + -0.07)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:56:06 2011