
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 21,114,709 – 21,114,800 |
| Length | 91 |
| Max. P | 0.692765 |

| Location | 21,114,709 – 21,114,800 |
|---|---|
| Length | 91 |
| Sequences | 12 |
| Columns | 114 |
| Reading direction | forward |
| Mean pairwise identity | 64.32 |
| Shannon entropy | 0.68265 |
| G+C content | 0.57366 |
| Mean single sequence MFE | -30.91 |
| Consensus MFE | -9.95 |
| Energy contribution | -9.36 |
| Covariance contribution | -0.59 |
| Combinations/Pair | 1.60 |
| Mean z-score | -1.61 |
| Structure conservation index | 0.32 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.43 |
| SVM RNA-class probability | 0.692765 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 21114709 91 + 23011544 ---ACAGUGCCACUCCGGCGCUGGCACAGCAAAUGAAUUUGGCCCGAAGAGCCUGCUGGAGGAUCAAAUCCUACAGCUCGGAAUCCCUCUUCAA-------------------- ---.(((((((.....)))))))((.(((.........)))))..((((((((.((((.(((((...))))).))))..)).....))))))..-------------------- ( -31.40, z-score = -1.73, R) >droSim1.chr2L 20694376 91 + 22036055 ---ACAGUGCCACUCCGGCGCUGCAACAACAGAUGAAUUUGGCCCGAAGAGCUUGCUGGAAGAUCAAAUCCUCCAGCUCGGAAUCCCUCUUCAA-------------------- ---.(((((((.....)))))))......((((....))))....((((((...((((((.(((...))).))))))..((...))))))))..-------------------- ( -29.50, z-score = -2.16, R) >droSec1.super_18 1033054 91 + 1342155 ---ACAGUGCCACUCCGGCGCUGCAACAACAGAUGAAUUUGGCCCGAAGAGCUUGCUGGAGGAUCAAAUCCUCCAGCUCGGAAUCCCUCUUCAA-------------------- ---.(((((((.....)))))))......((((....))))....((((((...((((((((((...))))))))))..((...))))))))..-------------------- ( -36.00, z-score = -3.58, R) >droYak2.chr2R 19484591 91 - 21139217 ---ACAGUGCCACUCCGGCGCUGCAACAGCAAAUGAAUUUGGCCCGAAGAGCCUGCUGGAGGAUCAAAUCCUCCAGCUCGGAAUCCCUCUUCAA-------------------- ---.(((((((.....)))))))......((((....))))....((((((((.((((((((((...))))))))))..)).....))))))..-------------------- ( -37.20, z-score = -3.81, R) >droEre2.scaffold_4845 19210775 91 - 22589142 ---ACAGUGCCACUCCGGCGCUGCAACGGGAAAUGAAUUUGGCCCGAAGAGCCUGCUGGAGGAUCAAUUCCUUCAGCUCGGAGUACCUCUUCAA-------------------- ---.(((((((.....)))))))....(((..(......)..)))((((((((.(((((((((.....)))))))))..)).....))))))..-------------------- ( -39.40, z-score = -3.63, R) >droAna3.scaffold_12943 1414381 79 - 5039921 ---ACAGUGCCACGCCGGCACUGGAACAGCAAAUGAAUCUGGCCAGGCGAGCCUGCUGGAGAA---------GCGGCAAGGACCUCCCCAC----------------------- ---.(((((((.....))))))).....((.......((..((.(((....)))))..))...---------...))..((......))..----------------------- ( -28.22, z-score = -0.67, R) >dp4.chr4_group1 307554 82 - 5278887 ---ACAGUGCCACGCCAGCACUGCAACAGCAAAUGAAUCUGGCCAGACGCGCCUGCUGGAGCA---------GCAGAAGGGAAGGUCCCUUGAA-------------------- ---...((((...(((((((.(((....)))..))...))))).....))))((((((...))---------))))(((((.....)))))...-------------------- ( -26.60, z-score = 0.01, R) >droPer1.super_5 246077 82 + 6813705 ---ACAGUGCCACGCCAGCACUGCAACAGCAAAUGAAUCUGGCCAGACGCGCCUGCUGGAGCA---------GCAGAAGGGAAGGUCCCUUGAA-------------------- ---...((((...(((((((.(((....)))..))...))))).....))))((((((...))---------))))(((((.....)))))...-------------------- ( -26.60, z-score = 0.01, R) >droWil1.scaffold_180772 4027384 87 - 8906247 ---ACAGUGCAACGCCAGCACUUGAACAACAAAUGAACCUUGCCCGCAGAGCCUGCUGGAGCAGGAGCCGCAAGAGCUCAACAAUUGAUG------------------------ ---..(((((.......))))).......(((.((....((((((((((...)))).)).))))(((((....).))))..)).)))...------------------------ ( -25.80, z-score = -1.39, R) >droVir3.scaffold_13246 475530 96 - 2672878 ---ACAGCGCGACGCCGGCGCUGCAACAACAAAUGAACCUCGCCCGGCGCGCCUGCUGGAGCAGCUGCCCGA---CCAGACCCAGCUCCAGUUCAAGUCCAA------------ ---.(((.(((.((((((.((....................))))))))))))))(((((((..(((.....---.))).....)))))))...........------------ ( -34.95, z-score = -1.45, R) >droMoj3.scaffold_6500 6032076 93 - 32352404 ---ACAGUGCGACGCCAGCGUUGCAGCAACAAAUGAACCUCGCCCGCCGCGCCUGCUGGAGCAGCUGCACAA---GCACAGC-AGCUCC--UCCCACUCCUA------------ ---.(((.(((..((..(((...((........)).....)))..))..))))))..((((.((((((....---.....))-)))).)--)))........------------ ( -29.30, z-score = -0.34, R) >droGri2.scaffold_15126 3760699 114 + 8399593 AUAGUGCUGCGACGCCGGCGUUGCAGCAACAAAUGAAUCUUGCCCGCCGCGCCUGCUGGAACAGCACCACAAAUCCCAAUAACAAUCCCAAUACAAAUCCAAGUCCGAGUCCGA ...((((((...(((.(((((.((.((((..........))))..)).))))).)).)...))))))............................................... ( -25.90, z-score = -0.62, R) >consensus ___ACAGUGCCACGCCGGCGCUGCAACAACAAAUGAAUCUGGCCCGAAGAGCCUGCUGGAGCAUCAAAUCCUACAGCACGGAAUCCCCCUUCAA____________________ ....((((((.......))))))...............(..((..(.....)..))..)....................................................... ( -9.95 = -9.36 + -0.59)

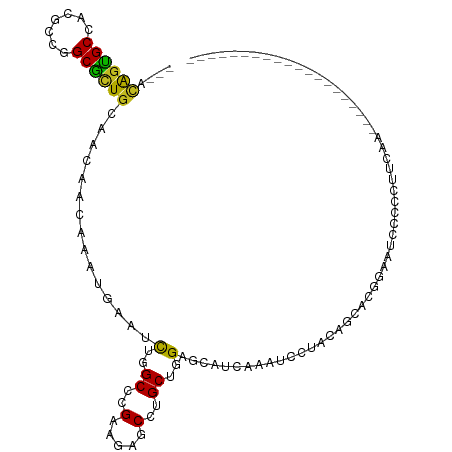

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:55:56 2011