
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 1,948,458 – 1,948,556 |
| Length | 98 |
| Max. P | 0.921015 |

| Location | 1,948,458 – 1,948,556 |
|---|---|
| Length | 98 |
| Sequences | 10 |
| Columns | 105 |
| Reading direction | forward |
| Mean pairwise identity | 62.87 |
| Shannon entropy | 0.80355 |
| G+C content | 0.37754 |
| Mean single sequence MFE | -20.47 |
| Consensus MFE | -8.66 |
| Energy contribution | -8.62 |
| Covariance contribution | -0.04 |
| Combinations/Pair | 1.50 |
| Mean z-score | -1.04 |
| Structure conservation index | 0.42 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.28 |
| SVM RNA-class probability | 0.921015 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 1948458 98 + 23011544 AUGCGAAAAACAA----AG-GCGAAAAUAUUAUG--CUUAUUUCGGGGGGGCUAUACAAAAAUUAUGGCGCCUGCGGAAAUUGUUUACGGCGGGUAAACAGUUUU .............----((-(((.........))--)))....((.(((.((((((.......)))))).))).))((((((((((((.....)))))))))))) ( -27.10, z-score = -2.60, R) >droEre2.scaffold_4929 1994383 77 + 26641161 AUGCUGGAAACAA----AGGCGGAAAAUAUUAUG--CUUAUUU----------------------UGGCGCCCGCGGAAAUUGUUUACGGCGGGUAAACAAUUUC .(((((....)).----.((((.((((((.....--..)))))----------------------)..)))).)))((((((((((((.....)))))))))))) ( -23.80, z-score = -2.60, R) >droYak2.chr2L 1929339 96 + 22324452 AUGCUAAAAACAA----AG-GCGAAAAUAUUAUG--CUUAUUUCGGGGG--CUGCACACAAAUUAUGGCGCCUGCGGAAAUUGUUUACGGCGGGUAAACAGUUUU .((((........----.)-)))...........--.......((.(((--(.((.((.......)))))))).))((((((((((((.....)))))))))))) ( -23.60, z-score = -0.93, R) >droSec1.super_14 1891691 97 + 2068291 AUGCUAAAAACAA----AG-GCGAAAAUAUUAUG--CUUAUUUCGGGGGG-CUAUAUAAAAAUUAUGGCGCCUGCGGAAAUUGUUUACGGCGGGUAAACAGUUUU .(((.........----..-.(((((.((.....--..)))))))(((.(-(((((.......)))))).))))))((((((((((((.....)))))))))))) ( -26.30, z-score = -2.59, R) >droSim1.chr2L 1915920 98 + 22036055 AUGCUAAAAACAA----AG-GCGAAAAUAUUAUG--CUUAUUUCGGGGGGGCUAUACAAAAAUUAUGGCGCCUGCGGAAAUUGUUUACGGCGGGUAAACAGUUUU .((((........----.)-)))...........--.......((.(((.((((((.......)))))).))).))((((((((((((.....)))))))))))) ( -27.30, z-score = -2.83, R) >dp4.chr4_group3 8709500 87 + 11692001 AUGCAAAAAAAUACAUCAGUUCGAAAUAGUUAUGGUCUGAUUUU------------------UGAUGGCGCCUGGGAAAAUUGUUUAUGGCGGGUAAACAGAUUU .(((.........((((((.(((...((....))...)))...)------------------))))).((((...(((.....)))..)))).)))......... ( -12.90, z-score = 1.03, R) >droPer1.super_1 10162182 86 + 10282868 AUGCAAAAAAAUACAUCAGUUCGAAAUAGUUAUGGUCUGAUUUU------------------UGAUGGCGCCUGGGGAAAUUGUUUAUGGCGGGUAAACAGAUU- .(((.........((((((.(((...((....))...)))...)------------------))))).((((...(((.....)))..)))).)))........- ( -12.50, z-score = 1.28, R) >droAna3.scaffold_12916 9658757 98 - 16180835 AUACUAAAAACAA----AACACAAAAUUAUUAUG--CUUAUUUUGAAGUGGGGGCA-AGAAAUUAUGGCGCCUCCGGAAAUUGUUUAUGGCGGGUAAACAAUUCA .............----....((((((.......--...))))))...(((((((.-............)))))))(((.((((((((.....))))))))))). ( -18.72, z-score = -0.60, R) >droWil1.scaffold_180772 1156127 103 - 8906247 AUUCAAAAUGAAAUAA-UAUUCAAAUACAAUGUA-CACAACAACAAAAAAUAUUAAGAACAGUUAUGGCGUGUUCACAAAUUGUUUACGCUGCAUAAACAAUUUG .(((.....))).(((-((((.........((..-..)).........))))))).(((((((....)).))))).(((((((((((.......))))))))))) ( -14.47, z-score = -0.60, R) >droGri2.scaffold_15252 96471 79 + 17193109 ----------------------AUGCACAAAACG-CUAGACAUUUGGAGCAAAUUUG--CAGCAAUGGCGCCUUUG-GAAUUGUUUAUGCGGUGUAAACAAUUUG ----------------------.....((((.((-(((.....(((..(((....))--)..))))))))..))))-(((((((((((.....))))))))))). ( -18.00, z-score = 0.08, R) >consensus AUGCUAAAAACAA____AG_GCGAAAAUAUUAUG__CUUAUUUCGGGGG__CU_UA_A_AAAUUAUGGCGCCUGCGGAAAUUGUUUACGGCGGGUAAACAGUUUU ............................................................................((((((((((((.....)))))))))))) ( -8.66 = -8.62 + -0.04)

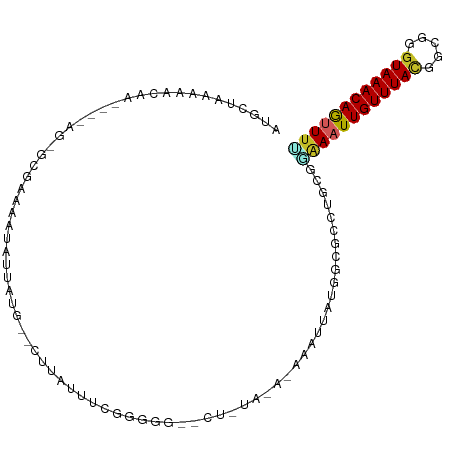

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:09:42 2011